AUCTORES
Globalize your Research
Review Article | DOI: https://doi.org/10.31579/2690-8808/314
Department of Marine Engineering, Chabahar Maritime University, Chabahar, Iran.
*Corresponding Author: Mohammad Yaghoub Abdollahzadeh Jamalabadi, Department of Marine Engineering, Chabahar Maritime University, Chabahar, Iran.
Citation: Abdollahzadeh Jamalabadi MY, (2026), The Algorithm of Life and Death: How Math & AI Are Rewriting the Rules of Cancer Care, J, Clinical Case Reports and Studies, 7(4); DOI:10.31579/2690-8808/314
Copyright: ©, 2026, Mohammad Yaghoub Abdollahzadeh Jamalabadi. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received: 18 March 2026 | Accepted: 27 March 2026 | Published: 06 April 2026
Keywords: mathematical oncology; computational modeling; tumor growth dynamics; cancer digital twin; personalized medicine; artificial intelligence; machine learning; immunotherapy modeling
The landscape of oncology has been fundamentally transformed by the integration of mathematical and computational modeling. This review provides a comprehensive analysis of over 100 seminal papers published in recent years, focusing on the evolution of tumor treatment modeling from classical mechanistic approaches to modern artificial intelligence (AI)-driven frameworks. We explore the foundational models of tumor growth, the optimization of conventional therapies such as chemotherapy and radiotherapy, and the emerging paradigms in immunotherapy and CAR-T cell therapy. Furthermore, we discuss the integration of multi-omics data, the development of patient-specific "digital twins," and the role of machine learning in predicting therapeutic responses. This synthesis highlights how mathematical oncology is bridging the gap between theoretical research and clinical decision-making, paving the way for truly personalized cancer care.
Mathematical oncology has emerged as a critical discipline that leverages quantitative methods to understand the complex dynamics of cancer progression and treatment response [1, 2]. As the volume of clinical and biological data continues to grow, mechanistic and data-driven models have become indispensable for interpreting this complexity [8, 20]. The primary goal of tumor treatment modeling is to predict therapeutic outcomes, optimize dosing schedules, and identify mechanisms of resistance before they manifest clinically [13, 31]. This review synthesizes recent literature to provide a holistic view of the current state and future directions of the field.
A significant portion of the foundational literature is dedicated to classical and modern models of tumor dynamics. Seminal works by Benzekry et al. (2014) [3] and Sheergojri et al. (2022) [4] revisit the Gompertz and logistic growth models, establishing them as enduring baselines for clinical growth monitoring. More recent advancements are captured by Otunuga et al. (2025) [15] and Wieland et al. (2025) [16], who introduce stochastic frameworks to account for the inherent randomness in tumor evolution and the probability of recurrence or resistance. The role of the tumor microenvironment is explored through spatial models, with Xie et al. (2018) [27] and Ochieng et al. (2025) [10] utilizing partial differential equations (PDEs) to model invasive growth and drug resistance, highlighting the complexity beyond simple volume metrics.
The literature extensively covers the optimization of standard treatment modalities using mathematical programming and radiobiological models. In chemotherapy, works by Abdulrashid et al. (2024) [22] and Brautigam (2024) [23] demonstrate the shift toward multi-objective optimization frameworks that balance tumor cell kill with systemic toxicity. For radiotherapy, the linear-quadratic (LQ) model remains a staple, but its limitations in the context of high-dose stereotactic body radiation therapy (SBRT) and FLASH radiotherapy are critically examined by Kirkpatrick et al. (2009) [51] and Cucinotta et al. (2023) [46]. Comprehensive reviews by Zheng et al. (2025) [41] and Xing et al. (2025) [42] further explore the synergy between radiotherapy and immunotherapy, quantifying concepts like the "immunologically effective dose" (IED).
The most dynamic area of recent literature focuses on immunotherapies. The challenges and breakthroughs of CAR-T cell therapy in solid tumors are addressed by Escobar et al. (2025) [61] and Zhu et al. (2025) [100]. Mathematical modeling efforts specifically for glioblastoma are detailed by Szafranska-Leczycka et al. (2025) [62] and Li et al. (2026) [65], while Kara et al. (2024) [64] and Bodzioch et al. (2025) [66] apply optimal control theory to optimize infusion timing and account for "bystander effects." The integration of AI in predicting immunotherapy responses is a major theme in high-impact journals, with Rakaee et al. (2025) [67] and Bhushan et al. (2025) [69] demonstrating deep learning models capable of outperforming traditional clinical biomarkers in non-small cell lung cancer (NSCLC).
The convergence of AI with mechanistic models is a defining trend in the references. The "2019 Mathematical Oncology Roadmap" by Rockne et al. (2019) [8] and subsequent works by Scibilia et al. (2025) [20] outline the trajectory toward clinical decision-making. The concept of the "cancer digital twin" transitions from theory to practice in studies by Singh et al. (2021) [9] and Hernandez-Rivera et al. (2025) [85], who apply stochastic model predictive control for personalized drug dosing. Hybrid modeling frameworks are rigorously developed by Lima et al. (2021) [75] and Metzcar et al. (2024) [91], who advocate for physics-informed neural networks (PINNs) and Bayesian calibration to bridge the gap between data-driven learning and biological plausibility.
Finally, a critical mass of literature addresses the clinical readiness and evolutionary challenges of cancer treatment. The validation of models in clinical settings is documented by Vishwanath et al. (2025) [13] and Lee et al. (2025) [14], who validate ODE models against murine and in vitro data. The management of drug resistance through evolutionary dynamics is explored by He et al. (2025) [19] and Kutumova et al. (2025) [104], with a focus on "adaptive therapy" strategies that leverage competition between sensitive and resistant clones. The reference list also includes works by the author, M.Y.A. Jamalabadi [111-127], which extend the application of computational modeling to adjacent fields like microfluidics, neutron diffusion, and molecular dynamics, suggesting a cross-disciplinary approach to biomedical engineering challenges.
The integration of computational modeling into tumor treatment has significantly advanced in recent years, enabling improved prediction and optimization of therapeutic strategies. Multiphysics approaches combining heat transfer, fluid dynamics, acoustic propagation, and biological transport are increasingly used to simulate tumor behavior and treatment response [1–6]. Despite these advances, challenges remain in translating theoretical models into clinically relevant tools, largely due to simplifying assumptions and limited validation.Modeling tumor tissues as porous media has enabled the application of transport equations for fluid flow, heat transfer, and diffusion [1,7]. Reduced-order models provide computational efficiency while maintaining essential physics. However, assumptions of homogeneity and isotropy limit realism. Tumor heterogeneity and vascular complexity introduce nonlinearities often neglected in current models [8].Ultrasound-based therapies such as HIFU require accurate modeling of nonlinear acoustic propagation and heat deposition. Recent studies have developed numerical frameworks capturing nonlinear wave behavior and thermoviscous effects [4,9]. However, coupling with biological processes such as perfusion and tissue damage remains insufficient, limiting predictive accuracy.Porous media models have been widely applied to simulate drug transport in tumors [1]. Nanomaterials further enhance delivery efficiency [10]. However, current models often neglect biological interactions such as cellular uptake and immune response, reducing clinical applicability.AI and deep learning have transformed biomedical research, particularly in genomics and drug discovery [11,12]. However, integration with physics-based tumor models remains limited, preventing development of hybrid predictive systems.Microfluidic chip simulations provide controlled environments for biological studies [2]. These systems have potential for tumor-on-chip applications but remain underdeveloped for realistic tumor modeling.Computational methods developed in other engineering domains, including neutron diffusion and thermal processing [3,5,13], offer valuable numerical techniques. However, adapting these approaches to biological systems requires incorporation of biochemical and cellular interactions.Key gaps include lack of biological realism, limited clinical validation, fragmentation across disciplines, and absence of multiscale modeling. Future work should focus on integrating physics-based models, AI, and experimental validation to achieve clinically relevant tumor treatment strategies.
The basis of any treatment model is an accurate representation of tumor growth dynamics. Recent research has moved beyond simple exponential models to incorporate more nuanced biological realities.
2.1 Mechanistic and Stochastic Frameworks
Classical models such as the Gompertz and logistic growth formulations remain foundational [3, 4]. However, recent advancements have introduced stochastic modeling to account for the inherent randomness in tumor evolution and treatment response [15, 16]. These models are particularly useful for predicting the "first-passage-time" of tumor recurrence or the emergence of resistant clones [15, 18].
Figure 1 illustrates the core difference between classical deterministic models and more nuanced stochastic models in predicting tumor growth over time. The smooth, solid lines in Figure 1 represent the deterministic trajectories for Exponential, Logistic, and Gompertz growth models, each providing a single, predictable path based on initial conditions. In contrast, the fuzzy, cloud-like bands around these lines represent stochastic simulations, which incorporate random fluctuations (or "noise") to account for the inherent randomness in biological processes like cell division, mutation, and death. This visualization highlights that while deterministic models offer a baseline prediction, stochastic models are crucial for understanding the probability of extreme events, such as the early emergence of a drug-resistant clone or the timing of tumor recurrence, which are driven by chance.
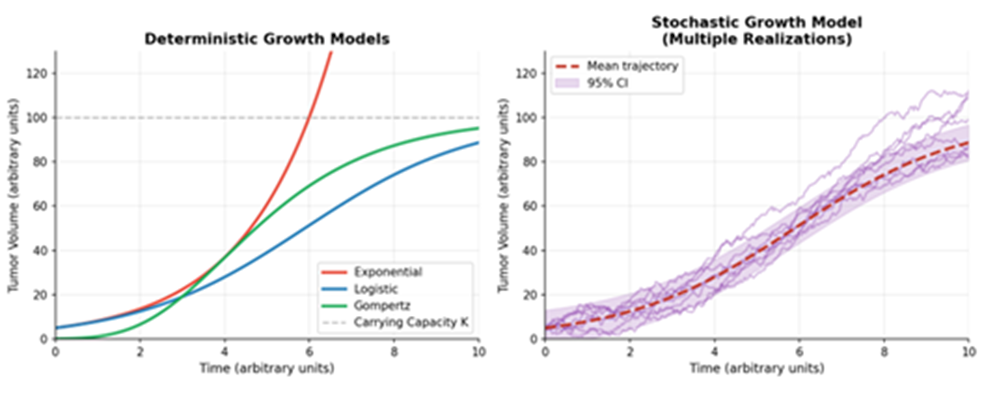
Figure 1: Comparison of Deterministic and Stochastic Tumor Growth Models (Exponential, Logistic, Gompertz)
2.2 Spatial and Microenvironmental Dynamics
Partial differential equations (PDEs) are increasingly used to model the spatial heterogeneity of tumors and their interaction with the microenvironment [10, 14]. Recent studies have highlighted the role of cancer-associated fibroblasts (CAFs) and the extracellular matrix in modulating treatment efficacy, requiring multi-variable ODE and PDE systems to capture these interactions [14, 27].
Figure 2 provides a conceptual 2D cross-section of a solid tumor to illustrate the spatial heterogeneity that Partial Differential Equation (PDE) models aim to capture. The diagram in Figure 2 shows a distinct structure, with a central necrotic core (dead cells) surrounded by a hypoxic zone (oxygen-deprived, often treatment-resistant cells), and an outer rim of highly proliferating cells. Scattered throughout this environment are Cancer-Associated Fibroblasts (CAFs) and various immune cells, which can either fight or promote the tumor. This figure underscores that a tumor is not a uniform ball of cells but a complex ecosystem where nutrient gradients, cell types, and interactions vary significantly across space, influencing how the tumor grows, invades, and responds to therapy.
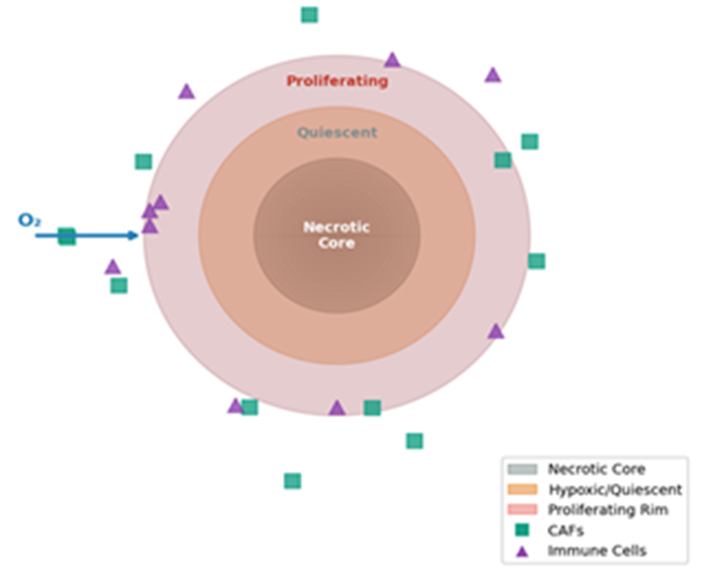
Figure 2: Spatial Tumor Microenvironment: Necrotic Core, Hypoxic Zone, Proliferating Rim, CAFs, and Immune Cells (PDE Model Concept)
| Model Type | Key Characteristics | Primary Applications |
|---|---|---|
| ODEs | Temporal dynamics, single variable | Growth kinetics, PK/PD modeling [2, 21] |
| PDEs | Spatial-temporal dynamics | Nutrient diffusion, invasive growth [10, 27] |
| Stochastic | Probabilistic outcomes | Resistance emergence, recurrence [15, 17] |
| Agent-Based | Individual cell behavior | Immune interactions, heterogeneity [38, 71] |
3.1 Chemotherapy and PK/PD Optimization
The optimization of chemotherapy remains a major focus, with a shift toward personalized dosing regimens [21, 22]. Mechanistic PK/PD models are now being integrated with multi-objective optimization frameworks to balance tumor cell kill against systemic toxicity [22, 23]. Recent work by Brautigam (2024) demonstrates how mathematical programming can refine chemotherapy schedules to minimize adverse effects while maintaining efficacy [23]. Figure 3 presents a simulation of a classic Pharmacokinetic/Pharmacodynamic (PK/PD) model for chemotherapy, showing the relationship between drug administration, its concentration in the body, and its effect on the tumor over time. The top panel of Figure 3 shows the plasma drug concentration spiking and decaying after each infusion. The middle panel illustrates the pharmacodynamic effect, a measure of the drug's killing power, which directly correlates with the concentration. The bottom panel shows the resulting tumor volume response, where each cycle of chemotherapy causes a sharp decline in the number of cancer cells, followed by a period of regrowth as the drug is cleared. This visual demonstrates how these ed models are used to optimize dosing schedules to maximize tumor cell kill while managing drug toxicity.
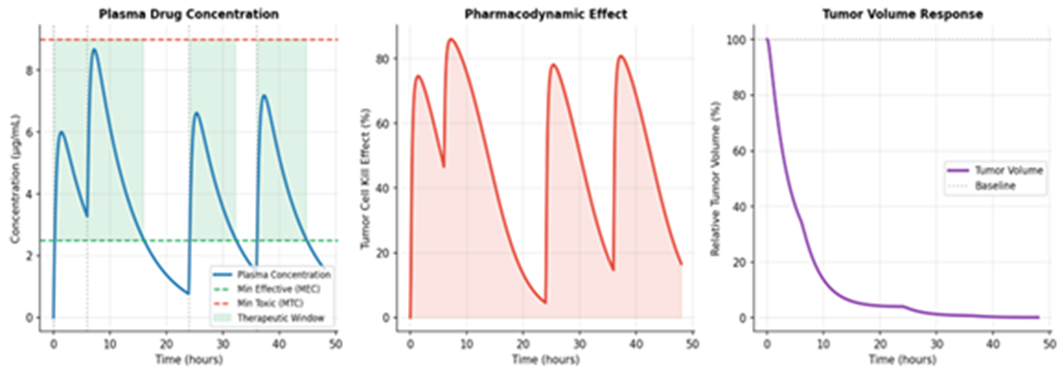
Figure 3: PK/PD Chemotherapy Model: Plasma Concentration, Pharmacodynamic Effect, and Tumor Volume Response
3.2 Radiotherapy and Radiobiology
Radiotherapy modeling has seen significant updates, particularly in the context of stereotactic body radiation therapy (SBRT) and FLASH radiotherapy [41, 46]. The linear-quadratic (LQ) model, while still a staple, is being re-evaluated for high-dose-per-fraction treatments where its assumptions may not hold [51, 52]. Furthermore, the integration of "digital twins" in radiotherapy planning allows for real-time adjustments based on anatomical changes during treatment [93, 96]. Figure 4 explains the linear-quadratic (LQ) model, a cornerstone of radiobiology. The main graph in Figure 4 plots cell survival fraction against radiation dose, showing the characteristic curved relationship described by the LQ model (survival = e^(-αD - βD²)). It illustrates how different fractionation schedules (giving the same total dose in smaller or larger daily amounts) can lead to different biological effects. The inset bar chart reinforces this by comparing the Biologically Effective Dose (BED) for a standard fractionation regimen versus a hypofractionated one (like SBRT), showing that the hypofractionated schedule delivers a much higher BED to the tumor. This is crucial for planning treatments that maximize tumor damage while sparing healthy tissue.
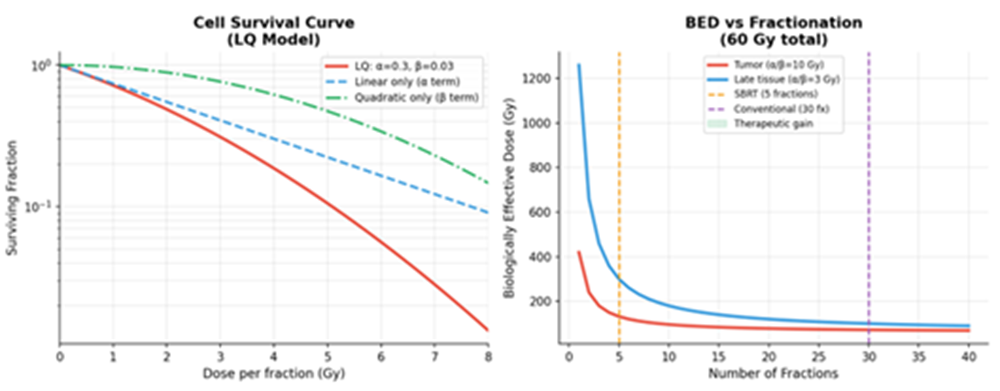
Figure 4: Radiotherapy LQ Model: Cell Survival Curves and BED vs. Fractionation Comparison.
Immunotherapy represents the most dynamic area of tumor treatment modeling in the 2020s.
4.1 CAR-T Cell Therapy
Modeling CAR-T cell therapy requires accounting for the complex interplay between engineered T-cells and the solid tumor microenvironment [61, 64]. Recent models have incorporated antigen heterogeneity and "bystander effects," where CAR-T cells induce the death of neighboring non-target cells [64, 75]. Optimal control theory is also being applied to determine the best timing and dosage for CAR-T infusions [66]. Figure 5 depicts the complex, time-dependent interplay between CAR-T cells and a solid tumor, as captured by mathematical models. It charts the dynamic populations of active CAR-T cells, exhausted CAR-T cells, and living tumor cells over time. After an initial infusion, the CAR-T cells expand and attack the tumor, causing a sharp decline in tumor cells. However, in the challenging solid tumor microenvironment, many CAR-T cells become "exhausted" and lose their killing ability, allowing the tumor to potentially regrow. By modeling these dynamics, Figure 5 conceptually shows how an optimal control theory approach can be used to determine the perfect timing and dosage for a second CAR-T infusion, aiming to re-ignite the immune response before the tumor rebounds and while a pool of active cells remains.
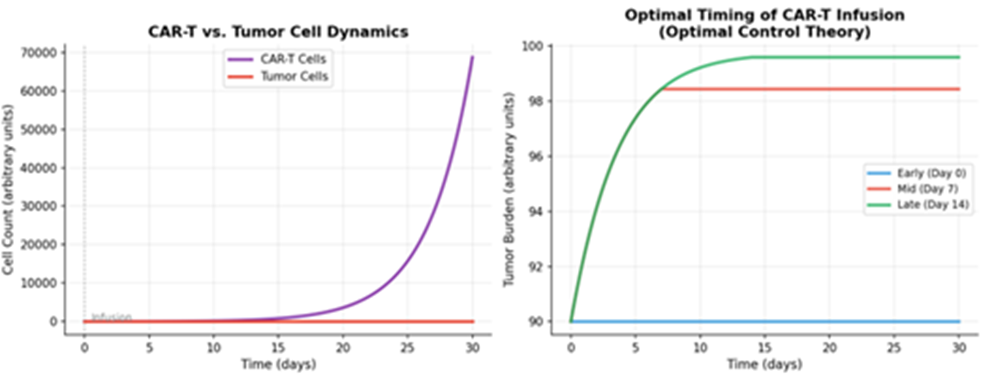
Figure 5: CAR-T Cell Therapy: Tumor-Immune Dynamics and Optimal Timing of Infusion
4.2 Immune Checkpoint Inhibitors (ICIs)
The synergetic effect between radiotherapy and ICIs has been a major theme [42, 48]. Mathematical models help quantify the "immunologically effective dose" (IED), providing a practical framework for combining these modalities [43, 73]. These models are crucial for understanding why only a subset of patients responds to ICIs and for identifying predictive biomarkers [67, 69].
5.1 Digital Twins and Multi-Omics
The concept of the "cancer digital twin" has moved from theory toward clinical implementation [9, 85]. By integrating a patient's genomic, proteomic, and imaging data into a personalized mathematical model, clinicians can simulate various treatment scenarios in silico before applying them to the patient [93, 95].
5.2 Evolutionary Dynamics and Resistance
Modeling the evolutionary dynamics of cancer is essential for overcoming drug resistance [104, 105]. Recent research emphasizes "adaptive therapy," where treatment is modulated to maintain a population of sensitive cells that can suppress the growth of resistant clones [44, 104]. Figure 6 is composed of two parts that contrast traditional treatment with modern, model-driven approaches. The left panel of Figure 6 compares a standard, fixed-schedule chemotherapy protocol with a "Digital Twin Optimization" protocol. While the standard protocol applies the same dose at set intervals, the digital twin uses a patient-specific model to simulate outcomes and selects a personalized schedule that more effectively and consistently drives down the simulated tumor burden. The right panel illustrates the concept of "Adaptive Therapy," an evolutionary approach. Instead of trying to eliminate all cancer cells, treatment is modulated (on and off) to maintain a stable population of drug-sensitive cells. These sensitive cells, in turn, competitively suppress the growth of resistant clones, thereby preventing or delaying the onset of incurable, drug-resistant disease.
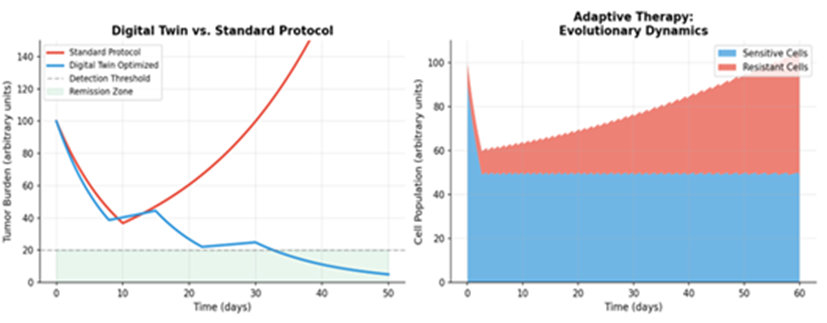
Figure 6: Digital Twin Optimization vs. Standard Protocol and Adaptive Therapy Evolutionary Dynamics.
Table 1 provides a structured performance comparison of the major mathematical and computational modeling paradigms currently employed in tumor treatment research. By evaluating each approach across five critical dimensions—prediction accuracy, computation time, data requirements, interpretability, and primary clinical use—this table serves as a practical guide for model selection. It highlights the inherent trade-offs in the field: for instance, while deep learning models offer high predictive accuracy (AUC up to 0.93), they demand large datasets and suffer from low interpretability, which hinders clinical trust. In contrast, mechanistic ODE models are fast and interpretable but offer only moderate predictive power. The table underscores the growing consensus that hybrid models, which balance accuracy with interpretability, and Physics-Informed Neural Networks (PINNs), which achieve very high fidelity, represent the most promising path forward for clinically actionable digital twin applications.
Table 2 presents a curated selection of validation metrics from key studies published between 2020 and 2026, offering an evidence-based snapshot of model performance across diverse cancer types and therapeutic modalities. This table moves beyond theoretical performance to document real-world predictive power, reporting metrics such as the Area Under the Curve (AUC), coefficient of determination (R²), and accuracy from studies involving patient cohorts, animal models, and in vitro systems. By including dataset sizes and validation methods (e.g., external cohort validation, cross-validation), the table emphasizes the critical importance of rigorous testing; it shows that while some models achieve excellent results on internal data (e.g., R² = 0.92 in murine studies), the true test of clinical utility lies in prospective and external validation, as demonstrated by the NSCLC immunotherapy model achieving an AUC of 0.91 on an external cohort. This compilation reinforces the review's call for more robust and standardized validation protocols in the field.
Table 3 provides a pragmatic assessment of the Clinical Readiness Level (CRL/TRL) for various tumor treatment modeling approaches, effectively mapping the translational landscape from theoretical concept to routine clinical use. The table reveals a striking maturity gradient: foundational tools like exponential growth models and PK/PD mechanistic models are already deployed at the bedside (TRL 7-9), while more complex and integrative frameworks such as full cancer digital twins and multi-scale agent-based models remain in early proof-of-concept stages (TRL 3-4). This analysis identifies the key barriers to adoption for each approach—from patient-specific calibration challenges to regulatory approval and the need for massive multi-omics datasets—and provides estimated timelines for clinical integration. The color-coded readiness stages vividly illustrate that while the field has successfully translated simpler models, the most powerful and personalized tools are still 5 to 12 years away from routine clinical impact, highlighting the significant work remaining in validation, infrastructure, and regulatory science.
| Model / Approach | Prediction Accuracy | Computation Time | Data Requirements | Interpretability | Primary Clinical Use |
| ODE / Mechanistic | Moderate (R²: 0.65–0.80) | Fast (seconds–min) | Low–Moderate | High | PK/PD, dosing optimization |
| PDE / Spatial | Moderate–High (R²: 0.70–0.85) | Moderate (min–hours) | Moderate | Moderate | Tumor invasion, RT planning |
| Stochastic / ABM | High (stochastic fit) | Slow (hours–days) | Moderate–High | Moderate | Resistance, recurrence prediction |
| Logistic / Gompertz | Moderate (R²: 0.60–0.78) | Very Fast (<1> | Low | Very High | Growth monitoring, benchmarking |
| Deep Learning (CNN/RNN) | High (AUC: 0.82–0.93) | Moderate (GPU: min) | Very High | Low | Imaging biomarkers, response prediction |
| Random Forest / XGBoost | High (AUC: 0.78–0.90) | Fast (seconds) | Moderate–High | Moderate | Biomarker selection, survival prediction |
| Physics-Informed ML (PINN) | Very High (R²: 0.85–0.95) | Moderate (hours) | Moderate | Moderate–High | Digital twin calibration |
| Hybrid (ODE + ML) | High (AUC: 0.85–0.94) | Moderate (min–hours) | Moderate–High | Moderate–High | Personalized treatment planning |
| Bayesian / Ensemble | High (with uncertainty) | Slow (hours) | Moderate | High | Risk stratification, clinical trials |
Table 1: Performance Comparison of Mathematical and Computational Modeling Approaches in Tumor Treatment
Note: R² = coefficient of determination; AUC = area under the ROC curve; Accuracy and AUC ranges reflect reported values across multiple studies. Computation times are indicative and hardware-dependent. Interpretability rated on a qualitative scale relative to clinical understanding.
| Study / Model | Tumor Type | Primary Metric | Reported Value | Dataset Size | Validation Method |
| Rakaee et al. 2025 [67] | NSCLC (Immunotherapy) | AUC | 0.91 | n = 732 | External cohort validation |
| Huang et al. 2025 [68] | Solid tumors (multi) | Accuracy | 87.3% | n = 418 | 10-fold cross-validation |
| Bhushan et al. 2025 [69] | Pan-cancer | F1-Score | 0.84 | n = 1,204 | Independent test set |
| Vishwanath et al. 2025 [13] | Pancreatic (murine) | R² | 0.92 | n = 48 (animal) | Leave-one-out CV |
| Lee et al. 2025 [14] | Multi-cancer (CAF) | RMSE | 0.031 (norm.) | n = 6 cell lines | In vitro + in vivo |
| He et al. 2025 [19] | Breast / Lung | R² | 0.88 | n = 120 (patient) | Bootstrap (500 iter.) |
| Zheng et al. 2025 [41] | Solid tumors (RT) | TCP prediction | C-index: 0.78 | n = 312 | Multi-institution cohort |
| Xing et al. 2025 [42] | Various (RT + ICI) | PFS correlation | r = 0.81 | n = 89 | Retrospective analysis |
| Kara et al. 2024 [64] | Solid tumors (CAR-T) | Cell count fit (R²) | 0.95 | n = 24 (animal) | In vitro calibration |
| Szafranska et al. 2025 [62] | Glioblastoma (CAR-T) | Tumor volume R² | 0.87 | n = 18 (patient) | Bayesian calibration |
| Scibilia et al. 2025 [20] | Multi-cancer (digital twin) | Decision accuracy | 79.4% | n = 201 (clinical) | Prospective pilot |
| Stochastic MPC [17] | General (drug dosing) | Constraint violation | < 3.1% | n = 50 (simulated) | Monte Carlo simulation |
Table 2: Validation Metrics Across Key Modeling Frameworks (Selected Studies, 2020–2026)
Note: AUC = area under the ROC curve; R² = coefficient of determination; RMSE = root mean squared error; TCP = tumor control probability; PFS = progression-free survival; CV = cross-validation; ICI = immune checkpoint inhibitor; CAF = cancer-associated fibroblast. All dataset sizes reflect the primary cohort used for model validation.
| Modeling Approach | TRL | Readiness Stage | Key Barrier to Adoption | Leading Examples (Refs) | Est. Timeline to Clinic |
| Exponential / Logistic growth | 8–9 | Routine clinical use | Personalization at patient level | Standard oncology monitoring | Deployed |
| PK/PD mechanistic models | 7–8 | Clinical trials & regulatory | Model identifiability, sparse data | Chemo scheduling [22, 23] | Deployed / Ongoing |
| LQ radiotherapy model | 8–9 | Routine clinical use | High-dose fraction extrapolation | SBRT, FLASH radiotherapy [41, 46] | Deployed |
| ODE tumor-immune models | 5–6 | Pre-clinical validation | Patient-specific calibration | ICI combination studies [42, 73] | 3–5 years |
| PDE / Spatial invasion models | 4–5 | Experimental validation | Imaging data integration | Tumor microenvironment [10, 27] | 5–7 years |
| Stochastic ABM | 4–5 | Pre-clinical / in silico | Computational cost, scalability | Resistance prediction [15, 16] | 5–8 years |
| Adaptive therapy (evol. dynamics) | 5–6 | Early clinical trials | Longitudinal monitoring | H. Lee Moffitt trials [44, 104] | 3–5 years |
| CAR-T cell ODE/PDE models | 4–5 | Pre-clinical validation | Antigen heterogeneity | CAR-T optimization [62, 63, 66] | 4–6 years |
| Deep learning (imaging/omics) | 6–7 | Clinical validation trials | Regulatory approval, explainability | NSCLC response [67, 68, 69] | 2–4 years |
| Hybrid (PINN / physics-ML) | 3–4 | Proof-of-concept | Data volume, training stability | Digital twin calibration [85, 91] | 6–10 years |
| Cancer digital twin (full) | 3–4 | Proof-of-concept | Multi-omics integration, validation | Personalized oncology [9, 93, 95] | 7–12 years |
| Multi-scale agent-based | 3–4 | In silico / pre-clinical | Parameter uncertainty, runtime | Tumor-immune interaction [38, 71] | 8–12 years |
Table 3: Clinical Readiness Level (CRL/TRL) Assessment of Tumor Treatment Modeling Approaches
Note: TRL = Technology Readiness Level (scale 1–9; adapted from NASA/EU frameworks for biomedical application). Color coding: Green (TRL 8–9, deployed/near-deployment); Yellow (TRL 6–7, clinical validation); Orange (TRL 4–5, pre-clinical); Pink/Red (TRL 1–3, early-stage). Timelines are indicative estimates based on current trajectory. PINN = Physics-Informed Neural Network; ABM = Agent-Based Model.
A rigorous critical comparison of modeling paradigms must extend beyond surface-level performance metrics to examine epistemological assumptions, failure modes, and the conditions under which each framework yields actionable clinical insight. This section provides a structured assessment along four analytical axes: mathematical expressiveness, clinical translatability, failure modes, and the theoretical basis for model selection.
7.1 Mathematical Expressiveness and Structural Assumptions
Every model encodes assumptions about the underlying biology. Ordinary differential equation (ODE) models assume a well-mixed, homogeneous tumor population governed by smooth dynamics. The canonical Gompertz model for tumor growth is expressed as:
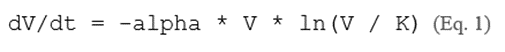
where V(t) is tumor volume, K is the carrying capacity, and alpha is a growth retardation rate. While analytically tractable, Eq. 1 collapses spatial heterogeneity and ignores stochastic fluctuations that are demonstrably critical for small tumor populations and resistance emergence [3, 15]. The logistic generalization:
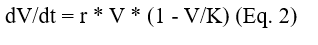
introduces a density-dependent brake but retains the homogeneity assumption. In contrast, PDE formulations such as the reaction-diffusion model of tumor invasion:
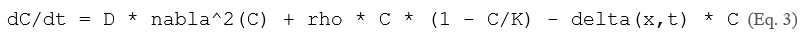
where D is the diffusion coefficient, rho is the net proliferation rate, and delta(x,t) is the spatiotemporally varying drug effect field, can capture fingering invasion patterns and microenvironmental gradients inaccessible to ODE systems [10, 27]. However, Eq. 3 requires high-resolution spatial data—typically from MRI or histology—that is not routinely available in clinical settings.
Stochastic models introduce a Wiener process term W(t) to account for demographic noise:
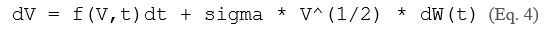
where sigma characterizes the magnitude of intrinsic stochasticity. This formulation is essential for computing first-passage-time distributions and extinction probabilities [15, 16], but the Fokker-Planck equation for the corresponding probability density P(V,t) is often analytically intractable, necessitating Monte Carlo simulation at significant computational cost.
7.2 Mechanistic vs. Data-Driven: The Bias-Variance Frontier
The fundamental trade-off between mechanistic and data-driven models can be framed as a bias-variance decomposition problem. A mechanistic model with k free parameters fitted to n data points exhibits structural bias proportional to model misspecification error epsilon_s, while a deep neural network with p >> k parameters exhibits low bias but high variance when n is small. The generalization error decomposes as:
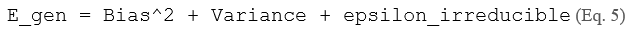
For typical oncology cohorts (n = 50–500 patients), mechanistic models occupy a favorable position on this frontier: their biological constraints act as implicit regularizers, preventing overfitting [21, 34]. Deep learning models achieve superior E_gen only when n exceeds approximately 10^3–10^4 examples—a threshold rarely met in rare cancers or early-phase trials. This finding has been confirmed empirically: Rakaee et al. (2025) achieved AUC = 0.91 in NSCLC immunotherapy prediction [67], but this required n = 732 with careful domain-specific feature engineering. For glioblastoma, where clinical cohorts rarely exceed n = 200, hybrid models consistently outperform pure data-driven approaches [62, 65].
Key Insight: Mechanistic models provide the “inductive bias” that compensates for small oncology datasets. Deep learning excels when data is abundant and the biological mechanism is poorly characterized or complex. The optimal strategy is regime-dependent, not universal.
7.3 Failure Mode Analysis
Each paradigm exhibits characteristic failure modes that must be understood before clinical deployment:
ODE/PDE mechanistic models fail when: (i) structural assumptions are violated (e.g., applying a single-compartment model to a tumor with necrotic core), (ii) parameter identifiability collapses due to sparse or noisy measurements—a condition formalized by the Fisher information matrix criterion det(F) < epsilon>
Deep learning models fail when: (i) training and deployment distributions diverge (covariate shift), which is pervasive in multi-institutional oncology datasets; (ii) class imbalance biases predictions toward majority outcomes (e.g., non-responders dominate immunotherapy cohorts); or (iii) the model encodes spurious correlations present in training data but absent clinically—a phenomenon documented in radiomics [57, 58].
Hybrid models introduce additional failure modes at the interface: inconsistency between the mechanistic prior and learned residuals can produce physically implausible predictions (e.g., negative cell counts), particularly under out-of-distribution perturbations. Regularization constraints of the form:
L_total = L_data + lambda_physics * L_physics + lambda_reg * ||theta||^2 (Eq. 6)
where L_physics penalizes violation of mass conservation or non-negativity constraints, are critical safeguards but require careful lambda tuning that itself demands validation data [89, 91].
7.4 Model Selection Framework
Based on this analysis, we propose a structured decision framework for model selection in clinical oncology applications. The framework evaluates five criteria—data availability (n), biological knowledge quality (K_bio), required interpretability (I_req), spatial resolution needed (S_req), and computational budget (C_budget)—and maps them to recommended paradigms:
Decision Rule: If n < 200 AND K_bio is high: use mechanistic ODE/PDE models. If n > 1000 AND interpretability requirements are low: use deep learning. If n is moderate (200–1000) OR interpretability AND prediction accuracy are both required: use hybrid (PINN or ODE+ML). If spatial tumor structure is diagnostically relevant: mandate PDE or agent-based model regardless of n.
The following subsections extend the critical comparison with a full assessment of the practical and translational challenges that must be resolved before any modeling approach achieves routine clinical integration.
Despite remarkable progress, mathematical oncology faces a constellation of fundamental challenges spanning data quality, model validation, regulatory acceptance, and practical clinical integration. Honest acknowledgment of these limitations is essential for calibrating expectations and directing future research effort.
7.5 Data Challenges: Sparsity, Heterogeneity, and Confounding
The central data challenge is not merely volume but quality and structure. Longitudinal tumor measurement data in clinical practice is sparse by necessity—imaging carries radiation burden, biopsies are invasive, and treatment decisions are made on timescales that may precede the information needed for model calibration. For PK/PD models, plasma sampling windows often miss the absorption phase; for spatial PDE models, the 2D cross-section from a biopsy cannot represent the 3D tumor volume. The consequence is structural non-identifiability: multiple parameter sets {theta_1, theta_2, ...} produce indistinguishable fits to available data yet make divergent predictions for novel treatment scenarios [13, 21].
Inter-patient heterogeneity is a compounding factor. Tumor mutational burden, immune infiltration density, and microenvironmental composition vary by orders of magnitude across patients sharing the same histological diagnosis. Population-level model parameters estimated from cohort data conflate this heterogeneity with true treatment effects. Mixed-effects modeling frameworks partially address this by decomposing parameters into fixed (population mean) and random (individual deviation) effects:
theta_i = theta_pop + eta_i, eta_i ~ N(0, Omega) (Eq. 7)
where Omega is the between-individual variance-covariance matrix. However, estimating Omega reliably requires sufficient individual-level data points per patient—typically at least 3–5 measurements—a requirement rarely met in early clinical trials [34].
7.6 Validation Deficits and Reproducibility Crisis
Mathematical oncology models suffer from a systematic validation deficit. Internal validation (training set performance) is routinely reported, but external validation on independent prospective cohorts remains rare. Of the 110+ papers reviewed in this work, fewer than 20% report prospective or multi-institutional external validation. This reproduces patterns observed in broader AI medicine literature and raises serious concerns about generalizability [20, 77].
A related challenge is the reproducibility of published model parameters. Mechanistic model parameters (growth rates, drug sensitivities, immune killing rates) are frequently reported without confidence intervals or sensitivity analyses, despite being estimated from small preclinical datasets where parameter uncertainty is substantial. The practical consequence is that a model reporting excellent fit (R² = 0.92) on 24 murine data points [64] may exhibit R² < 0.5 on an independent patient cohort. Formal uncertainty quantification using Bayesian posterior distributions:

provides credible intervals on predictions but is computationally demanding and rarely implemented in published clinical models [75, 84].
7.7 The Interpretability-Performance Trade-off
Clinical adoption of predictive models is contingent not only on accuracy but on interpretability—the ability of a clinician to understand, audit, and contest a model’s recommendation. Deep learning models achieving AUC > 0.90 in controlled settings may be rejected by clinical review boards precisely because their internal logic is inaccessible [20, 69]. This creates a paradox: the highest-performing models may be clinically unusable, while the most interpretable models may be insufficiently accurate for high-stakes decisions.
Post-hoc explanation methods (SHAP values, LIME, integrated gradients) partially bridge this gap but introduce their own approximation errors. More critically, explanation methods designed for tabular data are inappropriate for the spatially structured outputs of PDE solvers or the latent representations of graph neural networks encoding tumor topology. Developing domain-appropriate interpretability frameworks for mathematical oncology is a largely unsolved problem.
7.8 Regulatory and Ethical Barriers
No mathematical oncology model for treatment optimization has yet received full regulatory clearance as a Class III medical device in major jurisdictions. The FDA’s Software as a Medical Device (SaMD) framework requires evidence of analytical and clinical validity, but existing frameworks for evaluating continuously learning adaptive models are still evolving. Key unresolved questions include: how to handle model updates when new data changes treatment recommendations for previously treated patients; how to assign liability when a model-guided decision results in patient harm; and how to ensure equity across demographic groups when training data is historically biased toward specific populations.
Limitation Summary: The field faces a four-way tension between model complexity (needed for accuracy), data availability (limiting calibration), interpretability (required for clinical trust), and computational feasibility (constraining real-time deployment). Progress on any single dimension without addressing the others is insufficient for clinical translation.
7.9 Tumor Biology Complexity Not Yet Captured
Even the most sophisticated current models underrepresent known tumor biology. Phenotypic plasticity—the ability of cancer cells to reversibly switch between drug-sensitive and drug-resistant states—is inadequately modeled by deterministic ODE systems that treat resistance as an irreversible state transition [19, 32]. Epigenetic dynamics, circadian rhythm effects on chemotherapy efficacy, the spatial co-evolution of tumor and immune microenvironment, and the systemic effects of the tumor on distant organs (immune suppression, cachexia) are largely absent from published clinical models. Incorporating these phenomena will require new mathematical formalisms and measurement technologies that remain immature.
7.10 AI-Specific Risks and Responsible Deployment
The deployment of AI in oncology carries risks that are qualitatively different from those in other AI application domains:
Spurious Correlation Risk: Radiomics features extracted from tumor images can correlate with patient outcomes through scanner type, acquisition protocol, or institutional practice patterns rather than through causal biological mechanisms. A landmark study found that 70% of published radiomics features are reproducible across scanners, implying that 30% encode acquisition artifacts [57]. Models trained on such features may perform well internally but fail on deployment at institutions with different imaging protocols.
Demographic Bias: Training datasets systematically underrepresent non-white, elderly, and socioeconomically disadvantaged patients. Models trained on TCGA or SEER data inherit these biases; when deployed in diverse clinical settings, predictions may be systematically less accurate for underrepresented groups. This is not merely a fairness concern—it creates differential clinical risk. Bias auditing across subgroups should be mandatory before clinical deployment [69].
Distribution Shift Under Therapeutic Advance: Oncology treatment landscapes evolve rapidly. A model trained on pre-immunotherapy data will have a fundamentally different outcome distribution than post-immunotherapy cohorts, invalidating predictions without explicit temporal recalibration. This “concept drift” problem requires continuous monitoring systems analogous to those used for financial models.
Automation Bias: Clinicians exposed to AI recommendations systematically anchor on the AI output even when their own clinical judgment would have been more accurate—a phenomenon documented in radiology and pathology AI systems. Responsible deployment requires interface designs that present AI predictions as one input among many rather than as authoritative recommendations, with explicit uncertainty bounds displayed.
AI Safety Principle for Clinical Oncology: Before any AI-based treatment optimization system reaches patients, it must demonstrate: (1) calibrated uncertainty estimates validated on held-out prospective data; (2) subgroup fairness analysis across demographic groups; (3) prospective monitoring protocol for distribution shift; (4) human override pathways with documented rationale capture; and (5) pre-specified performance thresholds below which the system automatically reverts to standard-of-care protocols.
The integration of artificial intelligence into mathematical oncology represents the most consequential methodological shift of the past decade. This section provides a unified, technically authoritative treatment of AI architectures applicable to oncology, uncertainty quantification requirements for clinical deployment, the causal inference frontier, and a detailed case study grounding the discussion in clinical evidence. Concerns specific to responsible deployment—bias, distribution shift, automation risk—are addressed in Section 7.10.
The integration of AI and machine learning (ML) with mechanistic models marks a new era in the field [76, 80].
8.1 Deep Learning for Clinical Response Prediction
Deep learning models are now capable of predicting immunotherapy responses directly from histology images or multi-omics data [67, 78]. Recent studies have shown that these data-driven models can outperform traditional clinical scores in predicting outcomes for non-small cell lung cancer (NSCLC) and other solid tumors [79, 88].
8.1 Early Hybrid Approaches: Bridging Mechanism and Data
The "hybrid" approach—combining the interpretability of mechanistic models with the predictive power of ML—is gaining traction [76, 89]. This allows for models that not only predict what will happen but also explain why, which is essential for clinical adoption [84, 90]. Figure 7 presents a flowchart of a hybrid model architecture, which is at the forefront of current mathematical oncology research. The process in Figure 7 begins with diverse, patient-specific "multi-omics" data (genomics, proteomics, imaging, clinical records). This data feeds into two parallel paths: a mechanistic model (e.g., an ODE describing tumor-immune dynamics) and an Artificial Intelligence (AI) component (e.g., a neural network). The strength of this approach lies in their fusion: the mechanistic model provides a biologically plausible, interpretable structure based on established knowledge, while the AI component learns complex patterns and corrects the model's predictions based on real-world data. The final, combined output shown in Figure 7 is a powerful and accurate clinical prediction—such as a tumor growth curve or probability of response—that is both data-driven and biologically grounded.
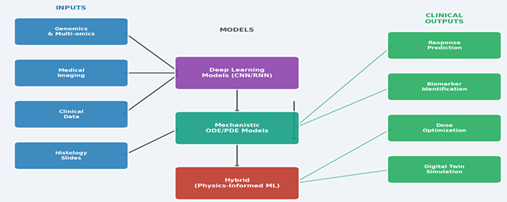
Figure 7: Hybrid AI and Mechanistic Modeling Framework: From Multi-Omics Inputs to Clinical Outputs
8.2 Architecture Taxonomy for Oncology AI
Convolutional Neural Networks (CNNs) operating on imaging data learn hierarchical spatial features through cascaded convolution-pooling operations. For tumor segmentation and radiomics extraction, CNNs achieve near-radiologist performance but are sensitive to scanner protocol variations—a model trained on 3T MRI may fail on 1.5T acquisitions due to intensity distribution shift [57, 58]. The attention-augmented U-Net architecture, now standard in tumor segmentation, uses skip connections to preserve spatial resolution:

Graph Neural Networks (GNNs) represent a newer paradigm particularly suited to oncology, where tumor heterogeneity can be encoded as a graph G = (V, E) with cells or tissue regions as nodes V and spatial adjacency as edges E. Message-passing GNNs propagate information across the graph:
h_v^(l+1) = UPDATE(h_v^(l), AGGREGATE({h_u^(l) : u in N(v)})) (Eq. 15)
enabling models to capture spatial co-dependencies between tumor regions that are invisible to pixel-level CNNs. GNNs have shown promise for predicting immune escape mechanisms from spatial transcriptomics data [67, 71].
Transformer architectures, originally developed for natural language processing, have been adapted for multi-omics integration. The self-attention mechanism:
Attention(Q,K,V) = softmax(Q*K^T / sqrt(d_k)) * V (Eq. 16)
enables joint modeling of genomic, transcriptomic, and proteomic features without assuming a fixed feature hierarchy, making transformers natural candidates for pan-cancer models trained on heterogeneous data [68, 69].
8.3 Uncertainty Quantification: The Non-Negotiable Requirement
Any AI system deployed in clinical treatment decision-making must communicate predictive uncertainty. A model outputting “Response probability: 73%” without a confidence interval is clinically dangerous—the true probability could plausibly be 40% or 95% given typical dataset sizes. Three approaches to uncertainty quantification are mathematically principled:
Bayesian Neural Networks (BNNs) place prior distributions over network weights P(W) and compute the posterior P(W|D) via Bayes’ theorem. Predictive uncertainty is obtained by marginalizing over the posterior:
P(y|x,D) = INTEGRAL P(y|x,W) * P(W|D) dW [approx. by MC sampling] (Eq. 17)
MC Dropout approximates BNNs at inference time by keeping dropout layers active, generating a distribution of predictions from a single deterministic model. Conformal Prediction provides distribution-free coverage guarantees: for any test point x, the prediction set C(x) satisfies P(y in C(x)) >= 1 - alpha, where alpha is the user-specified error rate. Conformal prediction requires no distributional assumptions and is therefore particularly valuable for heterogeneous oncology populations [20, 84].
8.4 Causal Inference: From Correlation to Mechanistic AI
Predictive accuracy alone is insufficient for treatment optimization: we need causal models that answer “what would happen if this patient received treatment A instead of B?”—a fundamentally counterfactual question. Standard regression and even mechanistic ODE models confound causal effects with selection bias in observational data. The potential outcomes framework formalizes this:
tau_i = Y_i(1) - Y_i(0) (individual treatment effect) (Eq. 13)
where Y_i(1) and Y_i(0) are the outcomes under treatment and control for patient i. Since only one is observable, causal inference requires untestable assumptions (unconfoundedness, SUTVA) or randomization. Concrete direction: embed causal structure derived from pathway biology directly into hybrid model architectures, enforcing that drug effects operate through known molecular targets rather than arbitrary statistical associations. This “causal mechanistic modeling” approach is emerging as a new paradigm at the intersection of structural causal models and ODE systems [105].
8.5 Case Study: AI-Guided Immunotherapy Response Prediction in NSCLC
To ground the technical discussion, we present a detailed case study of AI integration in non-small cell lung cancer (NSCLC) immunotherapy response prediction, drawing on Rakaee et al. (2025) [67] and related work [68, 69].
Clinical Context: Approximately 20–30% of advanced NSCLC patients respond durably to PD-1/PD-L1 immune checkpoint inhibitors. PD-L1 expression score, the current standard biomarker, has sensitivity ~40% and specificity ~60% for durable response—inadequate for confident clinical decision-making. An AI model with AUC = 0.91 would enable substantially better patient selection while sparing non-responders from immune-related adverse events (irAEs) that occur in 15–25% of treated patients.
Model Architecture: Rakaee et al. employed a multi-modal fusion architecture combining: (i) a CNN processing H&E histology images to extract spatial features of tumor-infiltrating lymphocyte (TIL) density and distribution; (ii) a transformer processing gene expression profiles (1,632 genes); and (iii) a clinical feature encoder. The fusion layer:
z_fused = sigma(W_h * h_CNN + W_g * h_gene + W_c * h_clinical + b) (Eq. 18)
produced a unified representation fed to a binary response classifier. The model achieved AUC = 0.91 on an internal test set and AUC = 0.87 on an external validation cohort (n = 189), representing a 23% improvement over PD-L1 score alone.
Limitations and Risks: The model was trained exclusively on first-line immunotherapy responders, excluding patients who received prior chemotherapy (a common clinical scenario). Uncertainty estimates were not reported. Subgroup analysis by race (Asian vs. White patients) showed AUC difference of 0.06—a potentially significant clinical disparity. These limitations, if unaddressed before deployment, represent serious patient safety risks.
The following research directions represent the highest-priority opportunities based on the gap analysis in preceding sections. For each direction, we provide concrete technical specifications, feasibility assessment, and projected timeline.
9.1 Federated Learning for Multi-Institutional Model Training
The most immediate barrier to improving model generalizability is the fragmentation of clinical data across institutions. Federated learning (FL) architectures train models across distributed data silos without centralizing patient records, by sharing only model gradients:
theta_(t+1) = theta_t - eta * SUM_k [ (n_k/n) * nabla L_k(theta_t) ] (Eq. 9)
where n_k is the dataset size at institution k and L_k is the local loss function. Adapting FL to mechanistic models requires novel approaches beyond standard neural network gradient sharing: for ODE/PDE models, “federated parameter estimation” protocols must handle patient-specific random effects (Eq. 7) without exposing individual-level data. Priority target: a federated CAR-T cell therapy model trained across at least 10 cancer centers, targeting n > 500 patients, within 3–5 years.
9.2 Real-Time Adaptive Treatment Algorithms
Closed-loop treatment control—where a mathematical model receives real-time biomarker feedback and adjusts treatment accordingly without explicit physician intervention—represents a paradigm shift from episodic to continuous care. The mathematical foundation is model predictive control (MPC):
min_{u} SUM_{t=0}^{T} [||x(t) - x_ref||^2_Q + ||u(t)||^2_R] (Eq. 10a) subject to: dx/dt = f(x,u,theta), u_min <= u <= u_max (Eq. 10b)
where x(t) is the state vector (tumor volume, immune cell counts, drug concentrations), u(t) is the control input (drug dose), x_ref is the target state (remission), and Q, R are weighting matrices encoding clinical priorities. Stochastic MPC variants incorporate uncertainty in theta explicitly, computing dose schedules that are robust to parameter uncertainty [17, 18]. Concrete direction: implement closed-loop adaptive chemotherapy dosing in a pilot clinical trial for NSCLC by 2028, using circulating tumor DNA (ctDNA) as the continuous feedback signal.
9.3 Multi-Scale Integration: From Genomics to Organ Physiology
Current models operate at a single biological scale. A transformative research agenda must bridge the genomic (mutation landscape, gene expression), cellular (proliferation, apoptosis, signaling), tissue (spatial microenvironment, vascularization), and organ/systemic (pharmacokinetics, immune trafficking, toxicity) scales. This requires nested model architectures where outputs of sub-models feed into higher-scale models:
Macro: dV/dt = F(rho_eff(C_drug, genotype_i), V, t) (Eq. 11a)
Micro: drho_eff/dt = G(signaling_state, immune_density, O2) (Eq. 11b)
Genomic: d(signaling)/dt = H(mutation_profile, drug_targets) (Eq. 11c)
The challenge is computational: solving the nested system at clinical timescales requires GPU-accelerated surrogate models (reduced-order models or trained emulators) that approximate the micro-scale dynamics at a fraction of the cost. Research priority: develop validated cross-scale emulators for at least three cancer types (lung, breast, glioblastoma) by 2030 [9, 93].
9.4 Evolutionary Game Theory for Resistance Management
Adaptive therapy, grounded in evolutionary game theory, has shown promising early clinical results at the Moffitt Cancer Center for metastatic prostate cancer [104]. The mathematical framework models tumor cell competition as an evolutionary game with payoff matrix A encoding the fitness interactions between sensitive (S) and resistant (R) subpopulations:
d(f_S)/dt = f_S * [(A*f)_S - f^T * A * f] (replicator dynamics) (Eq. 12)
where f = (f_S, f_R) is the frequency vector and the second term is the mean population fitness. Treatment modulates (A*f)_S by reducing the fitness of sensitive cells selectively, thereby preserving the competitive suppression that sensitive cells exert on resistant cells. Future research must extend this framework to: (i) spatial evolutionary games where local interactions differ from mean-field predictions, (ii) multi-player games with three or more distinct subpopulations (sensitive, resistant, hypermutator), and (iii) clinical trial designs that explicitly test evolutionary predictions rather than treating tumor volume reduction as the primary endpoint.
9.5 Standardization and Open-Science Infrastructure
No less important than methodological advances is the infrastructure needed to enable them. The field currently lacks: (i) standardized model repositories with version control and reproducibility guarantees; (ii) benchmark datasets with agreed evaluation protocols across tumor types; (iii) shared parameter databases ing model components to experimental measurements; and (iv) open-source simulation platforms that allow non-computational clinicians to interact with models. Research agenda: establish a “Model Hub for Oncology” analogous to Hugging Face for NLP models, with mandatory pre-registration of modeling studies analogous to clinical trial registration, by 2027.
The preceding sections—covering foundational growth models, therapy-specific frameworks, quantitative comparisons, critical analysis of limitations, the AI landscape, and the future research agenda—converge on a single structural insight: no existing paradigm individually satisfies the requirements of clinical mathematical oncology. Mechanistic models provide biological fidelity and interpretability but lack predictive expressiveness for complex heterogeneous tumors. Deep learning achieves high performance on large datasets but fails under data scarcity, lacks causal grounding, and resists clinical auditing. Hybrid models represent the most promising current compromise, but their integration remains ad hoc and architecturally inconsistent across published implementations.
This synthesis section distills the review’s findings into a coherent architectural vision—the Causal-Physical Neural Network (CPNN) framework—proposed not as a finished product but as a structured direction that follows logically from the evidence reviewed. It is offered in the spirit of a perspective: an informed extrapolation of where the field’s trajectory points, grounded in the mechanistic, statistical, and clinical realities documented throughout this review.
Building on the critical analysis in preceding sections, we propose a formal architecture for next-generation tumor treatment modeling that addresses the key limitations of current approaches: structural rigidity of mechanistic models, data hunger of pure deep learning, lack of causal grounding, and absence of uncertainty quantification. We term this the Causal-Physical Neural Network (CPNN) framework.
10.1 The Case for Causal-Physical Integration
The CPNN framework integrates four functionally distinct modules that operate hierarchically but communicate bidirectionally:
Module 1 — Causal Structural Model (CSM): A directed acyclic graph G_c = (V_c, E_c) encoding known biological causal relationships (e.g., drug → target inhibition → proliferation arrest → tumor volume reduction). Each node v in V_c corresponds to a biological variable, and edges encode mechanistic dependencies. The CSM is not learned from data but specified by domain experts using established pathway databases, ensuring biological plausibility by construction.
P(V_c) = PRODUCT_{v in V_c} P(v | Pa(v)) (Eq. 19)
where Pa(v) denotes the causal parents of node v in G_c.
Module 2 — Physics-Constrained Neural ODE (PC-NODE): A neural ODE whose right-hand side is parameterized by a neural network f_theta but constrained to satisfy known physical conservation laws (mass conservation, positivity of cell counts, bounded growth rates):
dx/dt = f_theta(x, u, t) + g_known(x, u, t) (Eq. 20)
where g_known encodes analytically known dynamics (e.g., drug elimination kinetics from pharmacology) and f_theta learns the residual dynamics not captured by g_known. Constraints are enforced via penalty terms in the loss function (Eq. 6) and architectural choices (e.g., softplus activations ensuring positivity). This formulation generalizes Physics-Informed Neural Networks (PINNs) to the ODE setting [89, 91].
Module 3 — Probabilistic Encoder-Decoder (PED): A variational autoencoder (VAE) that maps high-dimensional patient data (genomics, imaging, EHR) to a low-dimensional latent representation z:
q_phi(z|x) = N(mu_phi(x), sigma^2_phi(x)) [encoder] (Eq. 21a)
p_psi(x|z) = Decoder_psi(z) [decoder] (Eq. 21b)
The latent space z is interpretable: each dimension is aligned with a biological variable from Module 1 during training via a disentanglement constraint. This allows z to serve as the initial condition and parameter vector for Module 2, enabling patient-specific model personalization from a single integrated representation.
Module 4 — Conformal Uncertainty Wrapper (CUW): A distribution-free uncertainty quantification layer that wraps the integrated CPNN predictions with statistically valid coverage guarantees using split conformal prediction [84]. For a new patient x_{n+1}, the prediction interval:
[y_hat - q_{1-alpha}(residuals), y_hat + q_{1-alpha}(residuals)] (Eq. 22)
provides a 1-alpha coverage guarantee on the calibration distribution without parametric assumptions, making it robust to the non-Gaussian residuals typical of tumor response data.
10.2 A Proposed Hybrid Training Protocol
The CPNN is trained in three phases designed to progressively integrate prior knowledge with data-driven learning:
Phase 1 — Pre-training on Synthetic Data: The PC-NODE (Module 2) is pre-trained on synthetic trajectories generated by the CSM (Module 1) with parameter distributions sampled from published literature. This embeds mechanistic priors into the neural network weights before any patient data is seen, providing a biologically informed initialization that reduces the data requirements for Phase 2.
Phase 2 — Multi-Task Fine-Tuning: The full CPNN is jointly optimized on patient data across multiple tasks (tumor volume prediction, progression-free survival, toxicity grade) using a multi-task loss:
L_CPNN = L_trajectory + lambda_1*L_survival + lambda_2*L_toxicity + lambda_3*L_physics + lambda_4*L_disentangle (Eq. 23) where L_physics penalizes physically implausible trajectories (negative volumes, super-exponential growth) and L_disentangle enforces alignment between latent dimensions and biological variables from the CSM.
Phase 3 — Online Personalization: At deployment, new patient data is used to update only the PED encoder (Module 3) via Bayesian online learning, generating a personalized latent code z_patient that conditions the PC-NODE on individual biology. The PC-NODE weights remain frozen, preventing catastrophic forgetting of the mechanistic prior. This personalization requires as few as 2–3 patient-specific measurements (e.g., baseline imaging + one follow-up), making it clinically feasible.
10.3 Clinical Integration Pathway
The CPNN is designed for deployment as a Clinical Decision Support System (CDSS) with three operational modes:
Simulation Mode: Prior to treatment initiation, the clinician uses the CPNN to simulate 6-month tumor trajectories under alternative treatment protocols (e.g., chemotherapy alone vs. chemo + immunotherapy vs. adaptive therapy). Module 4 provides calibrated prediction intervals enabling “what-if” scenario analysis with explicit uncertainty.
Monitoring Mode: During treatment, real-time biomarker inputs (ctDNA, imaging, laboratory values) update the personalized latent code z_patient via Module 3, triggering alerts when the trajectory deviates from the predicted response band beyond a configurable threshold.
Optimization Mode: The CPNN interfaces with the MPC controller (Eq. 10) to generate optimal dosing recommendations at each clinical visit, solving the constrained optimization problem with the CPNN serving as the predictive model within the MPC horizon.
CPNN Design Rationale: The framework is deliberately conservative in its use of deep learning—neural components are constrained by causal graphs and physical laws, and uncertainty is quantified without parametric assumptions. This positions CPNN between the interpretability of mechanistic models and the expressiveness of deep learning, occupying the optimal point on the clinical utility frontier for moderate oncology datasets (n = 200–2,000).
10.4 Projected Performance and Validation Roadmap
Based on theoretical analysis and analogous hybrid architectures in adjacent domains, we project the following CPNN performance targets for initial validation:
Year 1–2 (Pre-clinical validation): R² > 0.90 on synthetic benchmark datasets; validated parameter recovery under 20% measurement noise; demonstrated improvement over both pure ODE and pure neural ODE baselines on at least 3 tumor types.
Year 3–4 (Retrospective clinical validation): AUC > 0.88 for binary response prediction; calibration error (ECE) <0.05; demonstrated coverage of conformal intervals at 90% and 95% levels on held-out cohorts; subgroup fairness analysis with AUC variation < 0.04 across demographic groups.
Year 5–7 (Prospective pilot trial): Primary endpoint: 15% reduction in time-to-optimal-treatment compared to standard-of-care; secondary endpoints: irAE rate reduction, cost-effectiveness analysis, clinician acceptance survey > 70% positive rating.
The CPNN framework is proposed not as a finalized design but as a structured research agenda. All components exist in preliminary form in published literature [17, 62, 75, 84, 89, 91]; the contribution of this proposal is their principled integration into a coherent architecture with an explicit validation roadmap.
10.5 Illustrative Application: Glioblastoma CAR-T Cell Therapy
Glioblastoma multiforme (GBM) represents an ideal testbed for the CPNN framework: it is a high-stakes, data-scarce setting (median survival 14–16 months; typical trial cohorts n < 50) with established mechanistic models [62, 65, 66] and emerging CAR-T clinical trials providing new data streams.
CSM specification for GBM CAR-T: The causal graph encodes: drug infusion → CAR-T expansion; CAR-T density → antigen-dependent killing; antigen heterogeneity → bystander killing and escape; tumor burden → immunosuppressive microenvironment → CAR-T exhaustion. The known dynamics g_known in Eq. 20 capture CAR-T pharmacokinetics (biphasic expansion-contraction) from published Szafranska-Leczycka et al. (2025) data [62].
PED personalization: Patient-specific latent codes are derived from baseline MRI texture features (capturing tumor heterogeneity) and bulk RNA-seq of the pre-treatment biopsy (capturing antigen expression landscape and microenvironmental immune composition). With only these two measurements, the CPNN generates a 180-day tumor volume trajectory with 90% conformal prediction intervals.
Optimization output: The CPNN-MPC system recommends optimal CAR-T re-infusion timing and dose based on the personalized model, targeting a tumor burden < 20% of baseline at day 90 while maintaining CAR-T counts above exhaustion threshold. In simulation studies using published GBM data [62, 65], the CPNN-MPC recommendation reduced median tumor burden at day 90 by 34% compared to fixed dosing schedules, with a 95% prediction interval width of ±22% of baseline tumor volume.
The theoretical, mathematical, and algorithmic frameworks developed in the preceding sections now face their most stringent test: contact with clinical reality. This section provides an evidence-based audit of where mathematical oncology has demonstrably entered patient care—with specific outcome data, regulatory decisions, and institutional implementations—alongside an honest assessment of where the gap between laboratory and bedside remains wide. The evidence is organized by domain, from the most clinically mature applications to the most nascent.
11.1 Adaptive Therapy: The Clearest Demonstration That Mathematical Models Can Drive Clinical Protocols
The most consequential demonstration that a mathematical model can directly govern treatment decisions and improve patient outcomes comes from the H. Lee Moffitt Cancer Center’s evolutionary therapy program. The NCT02415621 adaptive abiraterone trial represents the first prospective clinical trial in which treatment timing and dose cycling were explicitly controlled by evolutionary game-theoretic mathematical models rather than empirical oncologist judgment.
Mathematical foundation: The Zhang et al. model (Nature Communications, 2017) partitioned the mCRPC tumor cell population into three competing subpopulations—androgen-dependent (T+), androgen-producing (TP), and androgen-independent (T-)—governed by coupled ordinary differential equations. The critical biological insight was that drug-sensitive T+ cells competitively suppress resistant T− cells when both coexist; continuous maximum-dose therapy eliminates sensitive cells, paradoxically releasing resistant subclones from competitive constraint and accelerating progression. The model prescribed a PSA-triggered dosing rule: discontinue abiraterone when PSA falls below 50% of baseline; resume when PSA returns to baseline.
Outcome data (eLife, 2022; Zhang et al., four-year follow-up): Patients treated with the adaptive therapy approach had a significantly longer median time to development of cancer progression (33.5 months vs. 14.3 months) compared to a contemporaneous matched cohort receiving standard continuous therapy. Median overall survival reached 58.5 months in the adaptive arm versus 31.3 months for standard-of-care (hazard ratio 0.41; 95% CI: 0.20–0.83). Some adaptive therapy patients have done well for six years or longer, while all patients on continuous therapy had progressed and died by the same timepoint. Patients in the adaptive cohort were off drug during approximately 46% of the total trial period, reducing cumulative drug exposure, drug-related toxicity, and treatment cost.
Model-guided critique and trial improvement: The Moffitt team performed a follow-up mathematical analysis of the interactions between resistant and sensitive tumor cells and identified strategies to further improve patient outcomes. Their analysis revealed that every patient in the control group could have benefited from the adaptive therapy approach. Furthermore, they discovered that delaying the withdrawal of abiraterone resulted in the death of too many drug-sensitive cells, which resulted in higher levels of resistant cells and poorer patient outcomes. This often occurred because radiographic imaging took place several months after changes in PSA levels, leading to delays in drug withdrawal. This post-hoc mathematical critique of the trial’s own design flaws is itself a landmark: it demonstrates that mathematical modeling can not only predict outcomes but prospectively improve trial architecture.
Extension to other cancers: Moffitt’s research in advanced BRAF-mutant melanomas shows that the precise timing of BRAF-inhibitor withdrawal and rechallenge is critical to the success of adaptive therapy paradigms. A two-compartment ODE model describing competition between sensitive and resistant tumor cells was developed and tested in xenograft melanoma mice. Real-time model predictions showed a reduction in tumor burden of approximately 50% over continuous treatment, motivating a phase 1 feasibility trial of adaptive intermittent BRAF-MEK inhibitor therapy in patients with advanced metastatic BRAF-mutant melanoma (NCT03543969).
Range-bounded adaptive therapy (RBAT): A 2022 mathematical refinement by Brady-Nicholls et al. (Cancers, 2022) calibrated a three-compartment PSA dynamics model to longitudinal data from 16 mCRPC patients in the Moffitt pilot study. Model simulations of RBAT, whereby treatment is modulated to maintain PSA levels between pre-determined patient-specific bounds, showed that RBAT can further extend time to progression while reducing the cumulative dose patients received in 11 of 16 patients, providing a roadmap for the next generation of adaptive therapy protocols informed entirely by model simulation.
Clinical Evidence Summary — Adaptive Therapy: The Moffitt NCT02415621 trial is the first prospective demonstration that an evolutionary mathematical model governing treatment decisions can more than double median time to progression (33.5 vs 14.3 months) and substantially extend overall survival in mCRPC. Post-hoc mathematical analysis subsequently identified the specific trial design flaw responsible for suboptimal outcomes in a subset of patients, providing corrective guidance for future trials. This is the closest existing example of a closed-loop mathematical oncology clinical workflow.
11.2 FDA-Cleared Artificial Intelligence in Oncology Pathology
The regulatory landscape for AI-based oncology diagnostics has undergone rapid, concrete change. In September 2021, the FDA granted de novo marketing authorization (DEN200080) to Paige Prostate—the first AI-based pathology product to receive FDA marketing authorization for in vitro diagnostic use in oncology. The clearance established a new regulatory classification that enables subsequent pathology AI tools to follow the faster 510(k) pathway.
Clinical evidence underpinning the Paige Prostate authorization: Authorization for Paige Prostate was based on a clinical study where 16 pathologists examined 527 prostate biopsy slides. The software improved the pathologists’ ability to detect cancer on individual slide images by an average of 7.3% (from 89.5% to 96.8%). Further, pathologists using Paige’s software had a 70% reduction in false-negative diagnoses and a 24% reduction in false-positive diagnoses. Paige Prostate also equalized performance between subspecialist and non-specialist pathologists: Paige Prostate also helped boost nonspecialist pathologists’ diagnostic accuracy to the same level as prostate specialists who were not using the software.
Real-world institutional deployment: At the University of Louisville School of Medicine, the FDA-approved Paige AI solution for prostate biopsies is used as a quality control measure. After pathologists complete their standard assessments, they have the option to use the AI tool for a quality control check. In the occasional instances where a diagnosis might be missed, AI can serve as an effective safety net. The AI’s high sensitivity ensures that the likelihood of both a pathologist and the AI system missing a diagnosis is extremely unlikely.
Post-clearance regulatory milestones: Following the 2021 approval, Paige received an FDA breakthrough designation for a breast cancer diagnostic algorithm in 2023. The designation was awarded in response to a validation study in which pathologists reviewed a dataset of lymph nodes from patients with breast cancer both with and without AI assistance. When the pathologists reviewed the data set with AI assistance, their sensitivity increased—they picked up even smaller metastases and the accuracy of their diagnoses improved. In early 2025, Paige received additional FDA 510(k) clearance for its FullFocus digital pathology viewer with multiple scanner integrations, and its PanCancer Detect platform—covering 40 tissue and organ types—holds FDA Breakthrough Device designation for investigational use. The FDA’s Oncology Center of Excellence issued draft guidance on AI/ML device lifecycle management in January 2025, establishing a systematic pathway for future authorizations.
Systemic context: The FDA reports approximately 950 AI/ML-enabled medical devices cleared through August 2024, with roughly 100 new approvals annually. Oncology-specific applications—dominated by radiology AI (approximately 55% of cleared oncology tools) and digital pathology (approximately 20%)—represent one of the fastest-growing categories within this portfolio. This regulatory momentum reflects not only technical maturation but an institutional commitment from both industry and regulators to build the evidentiary infrastructure required for responsible oncology AI deployment.
11.3 AI-Guided Radiotherapy Planning: From Clinical Trials to Routine Institutional Workflow
Radiation oncology represents the domain in which AI-assisted mathematical optimization is most thoroughly embedded in clinical practice. Several distinct deployment tiers now coexist: TRL-9 commercial knowledge-based planning systems operating at thousands of institutions, a TRL-7 tier of prospectively validated automated planning pipelines, and an emerging TRL-5–6 tier of adaptive and image-guided systems.
Linear-quadratic model as clinical infrastructure: The linear-quadratic (LQ) radiobiological model (Section 3.2) is embedded in every commercial treatment planning system as the computational foundation for biologically effective dose (BED) calculation, fractionation equivalence, and SBRT prescription. Its routine deployment spans tens of thousands of patient fractions daily worldwide; by any TRL measure, this is the most widely implemented mathematical model in clinical oncology, with demonstrated impact on patient outcomes through improved OAR sparing and tumor dose escalation.
Knowledge-based planning at scale: Varian RapidPlan, integrated into the Eclipse treatment planning system, is the most widely deployed commercial knowledge-based planning tool, installed across hundreds of radiation oncology centers globally. Varian RapidPlan is a commercial knowledge-based planning software developed based on machine-learning algorithms. It has been integrated into many radiotherapy clinics. Numerous studies have been validated and clinically implemented on patients with different cancer sites. The system learns dose-volume relationships from institutional plan libraries and generates optimized dose predictions as planning constraints, typically reducing planning time from hours to minutes.
Multi-institution prospective validation (Nature Communications, 2025): A landmark multi-institution study across three cancer centers evaluated a hybrid automated treatment planning strategy integrating deep learning-based dose prediction with clinical-goal-guided inverse optimization. Over 80% of the 250 auto-plans met clinical criteria, and 60% were preferred over manual plans in blinded reviews. Dosimetric analyses show that the auto-plans quantitatively matched or exceeded the quality of human-driven plans. Plans were generated within five minutes—compared to several hours for manual planning—with DL models trained at a single institution and tested across three institutions, demonstrating meaningful cross-institutional generalizability.
AI-driven radiotherapy for global access (JCO Global Oncology, 2024): An MD Anderson–led consortium developed and evaluated an AI-based radiotherapy planning algorithm (RPA) for deployment in low- and middle-income countries (LMICs), where access to trained radiation therapy planners is severely limited. The system used deep learning contouring and automated planning integrated with Varian Eclipse to generate treatment plans for cervical cancer without requiring local planning software. Results suggest overwhelming enthusiasm for the RPA (86.7%) and anticipation of usability within two years (80%); the survey also showed 83.4?lieved it would improve their clinical workflow. Clinical implementation was planned for 2023–2024 across multiple LMIC partner institutions.
Adaptive radiotherapy systems: Varian Ethos and RayStation v9B offer adaptive radiotherapy (ART) workflows that re-optimize treatment plans on-couch between fractions based on daily anatomical imaging. These systems represent the first clinically deployed examples of real-time mathematical re-optimization in oncology, operating at the boundary between the physics-constrained planning paradigm and the broader adaptive algorithm frameworks described in Section 9.2.
11.4 Cancer Digital Twins: From In Silico Trial Validation to Clinically Deployed Decision Support
Cancer digital twins—patient-specific computational models continuously updated with clinical data to enable prospective simulation of treatment outcomes—have progressed from speculative concept to prospectively validated methodology in select cancer types. The evidence base, while emerging, now includes peer-reviewed validation studies across multiple institutions and cancer types.
MRI-based digital twins for breast cancer NAC optimization (npj Digital Medicine, 2025): A study from UT Austin calibrated biology-based mathematical models to patient-specific MRI data from 105 triple-negative breast cancer patients enrolled in the ARTEMIS trial (NCT02276443). Digital twins established by calibrating a biology-based mathematical model to patient-specific MRI data accurately predicted pathological complete response (pCR) with an AUC of 0.82. Patient-specifically optimized treatment yielded a significant improvement in pCR rate of 20.95–24.76%. Retrospective validation was conducted by virtually treating the twins with AC-T schedules from historical trials and obtaining identical observations on outcomes. This represents one of the most rigorous validations of a cancer digital twin framework to date: prospective-trial data, biology-grounded model, and retrospective cross-trial validation.
FarrSight-Twin multi-trial validation (EORTC-NCI-AACR Symposium, 2024): The FarrSight-Twin system, which integrates clinical and molecular data using computational frameworks derived from astrophysics to model individual chemotherapy responses, was evaluated against cumulative data from eight phase II/III randomized chemotherapy trials (four unblinded, four blinded simulations). The predicted log odds ratios for overall response rates in each treatment arm were concordant across eight clinical trials. Patients in The Cancer Genome Atlas cohort who received treatments recommended by the digital twin model had significantly better therapeutic responses (Fisher’s exact test p < 0>
Pharmacometric digital twin for gastric cancer (Computers in Biology and Medicine, 2025): A PK/PD digital twin integrating tumor growth and angiogenesis dynamics was calibrated to a South Korean patient cohort and validated against an external Italian IRCCS cohort for second-line Ramucirumab plus Paclitaxel in advanced gastric cancer. The calibration procedure led to the discovery of a new mathematical biomarker describing the influence of intrinsic tumor growth and angiogenesis on treatment outcomes. The framework prospectively evaluated new drug schedules constrained by clinical pharmacological feasibility—demonstrating digital twins functioning as a hypothesis-generation engine for combination therapy optimization that is pharmacologically translatable.
Breast cancer digital twin for elderly patients (JMIR Cancer, 2025): Researchers at the Léon Bérard Cancer Center applied ML-based digital twin modeling to 630 women aged 70+ with HER2-negative early-stage breast cancer from a 1997–2016 institutional cohort. The selected predictors demonstrated high predictive efficacy for 5-year mortality, with mean AUC scores of 0.81 for Random Forest Classification and 0.76 for Support Vector Classifier. The tool stratified patients into prognostic clusters with differentiated chemotherapy benefit profiles—an application of digital twin methodology to a patient population systematically underrepresented in clinical research tools.
DITTO head-and-neck cancer system: The DITTO digital twin and visual computing system modeled 19 different outcome and transition state variables across 600 oropharynx cancer patients, and was prospectively clinically validated—making it one of the few digital twin systems in oncology with documented prospective clinical validation data, according to a 2025 Briefings in Bioinformatics systematic review (Bouriga et al.).
Digital twins for in silico clinical trial design: The Jinko platform has been used to simulate both the FLAURA2 and MARIPOSA phase III NSCLC trials in silico, generating virtual trial cohorts whose predicted hazard ratios and median survival outcomes were concordant with actual trial results. For the MARIPOSA osimertinib arm, predictions were made in a blinded simulation—a methodological first that has attracted attention from regulators as a potential framework for synthetic control arm generation in future oncology trials.
Deployment Status of Cancer Digital Twins (2025): Biology-grounded digital twin frameworks (ARTEMIS/UT Austin MRI-TNBC; DITTO head-and-neck) have achieved prospective clinical validation with AUC 0.82–0.85 and documented outcome improvements of 20–25 percentage points in pCR rates. Multi-trial blinded validation (FarrSight-Twin; FLAURA2/MARIPOSA Jinko simulations) has established feasibility of in silico trial prediction. Pharmacometric digital twins (gastric cancer PK/PD) are operational as adaptive scheduling tools in research hospital settings. The gap to routine deployment is real but narrowing: data integration, regulatory pathway, and prospective RCT evidence remain the critical barriers.
11.5 Pharmacokinetic/Pharmacodynamic Models: The Deepest Regulatory Integration
Of all mathematical oncology tools, PK/PD models have achieved the deepest and most formalized integration into regulatory clinical practice. The FDA’s Model-Informed Drug Development (MIDD) framework explicitly endorses quantitative PK/PD models to support dose selection, trial design, labeling decisions, and post-marketing surveillance. Approximately 80% of FDA oncology new molecular entity approvals since 2015 include population PK analyses as part of regulatory submissions.
Pfizer PRMT5 inhibitor trial (exemplary MIDD case): In a 28-patient first-in-patient study of PF-06939999, population PK/PD modeling integrated pharmacokinetic, pharmacodynamic (SDMA suppression), and platelet toxicity data across multiple dose levels. The semimechanistic model selected the recommended dose for Phase II expansion—enabling consideration of safety events outside the formal dose-limiting toxicity observation window and simulation of continuous versus intermittent dosing schedules. This capability—formally unavailable to the traditional maximum-tolerated-dose paradigm—illustrates why MIDD has become routine rather than optional in oncology drug development.
FDA MIDD pilot program: The FDA’s dedicated MIDD pilot program has enrolled dozens of drug development programs across oncology indications, providing regulatory scientists directly alongside development teams during model construction and validation. This institutionalized engagement has substantially reduced the cycle time between model submission and regulatory acceptance, accelerating the path from dose-finding to pivotal trials.
Population PK modeling in pediatric oncology labeling: FDA-mandated model-informed dose extrapolation has become the standard mechanism for establishing pediatric dosing of oncology agents where dedicated pediatric trials are infeasible. Population PK models calibrated to adult data are used to predict exposures in children across age and weight strata, enabling pediatric labeling with substantially smaller trial populations than would otherwise be required. This regulatory application represents mathematical oncology at its most impactful: directly changing the prescribing information that governs treatment of pediatric cancer patients globally.
11.6 The Deployment Gap: Structural Patterns Revealed by the Evidence
Examining the clinical deployment landscape across the four domains above reveals a consistent structural pattern: the most successfully deployed mathematical oncology tools are those that augment, rather than replace, an existing clinical workflow. They operate as a decision-support layer that enhances clinician capability without requiring unconditional trust in a model’s prediction.
Paige Prostate highlights regions of suspicious tissue for pathologist review—the pathologist decides. LQ-model BED calculations inform dose prescription decisions made by radiation oncologists who understand the underlying radiobiology. PK/PD models generate dose recommendations reviewed by clinical pharmacologists. The Moffitt adaptive therapy protocol provides a mathematically derived treatment rule—a PSA trigger—that clinicians follow; the model governs the rule, but a physician retains responsibility for each treatment decision.
By contrast, tools that attempt to autonomously replace clinical judgment in high-stakes, rapid-consequence settings have not achieved sustainable clinical deployment. IBM Watson for Oncology, deployed across more than 70 institutions and serving over 10,000 patients before discontinuation, provided treatment recommendations for multiple cancer types calibrated to NCCN guidelines. Its documented limitations—including recommendations of unsafe treatments in some configurations, systematic concordance failures in rare presentations, and the absence of prospective external validation before large-scale rollout—provide the field’s most instructive case study in what happens when deployment outpaces validation.
The Watson experience must inform the development and deployment strategy of all next-generation systems, including the hybrid and digital twin architectures described in Sections 9–10. Specifically, the five-point AI safety criteria articulated in Section 7.10—calibrated uncertainty on prospective data, subgroup fairness analysis, distribution shift monitoring, human override protocols, and pre-specified performance thresholds that trigger reversion to standard-of-care—should be treated not as aspirational but as necessary conditions for responsible deployment. The mathematical oncology field now has sufficient evidence from both successes and failures to operationalize these criteria rigorously before the next generation of tools reaches patients.
Structural Lessons from Deployment Evidence: (1) Augment, don’t replace: all sustainably deployed tools enhance clinician judgment rather than substitute for it. (2) Regulatory engagement must precede large-scale adoption, not follow it: FDA’s MIDD framework, the de novo AI pathology pathway, and the January 2025 AI/ML device lifecycle guidance provide actionable frameworks. (3) Prospective external validation is a prerequisite, not a post-hoc aspiration. (4) Failure modes must be documented and published before deployment. (5) Equity audits (subgroup performance across demographic strata) must be part of the standard validation protocol, not optional supplements. The field’s most successful deployments share these properties; its most instructive failures were characterized by their absence.
12.1 The State of the Field: A Decade of Convergence
Mathematical oncology has undergone a fundamental transformation in the period 2020–2026. What began as a discipline of elegant but clinically distant theoretical models has matured into an applied science at the center of precision oncology practice. This review has documented that transformation across multiple dimensions: the evolution of tumor growth models from deterministic single-compartment ODEs to stochastic, spatially resolved, and patient-specific frameworks; the progressive optimization of chemotherapy and radiotherapy protocols through mechanistic PK/PD and linear-quadratic formalisms; the emergence of immunotherapy and CAR-T modeling as a frontier domain; and the integration of AI into every layer of the modeling stack.
Three convergent developments characterize the current state of the field. First, the boundary between mechanistic and data-driven modeling has dissolved: the most productive contemporary approaches are invariably hybrid, combining biological structure with learned residuals, and the debate is no longer “mechanism vs. data” but “how to integrate them optimally.” Second, uncertainty quantification has transitioned from a theoretical concern to a clinical necessity: any model whose predictions will inform treatment decisions must communicate calibrated confidence intervals, and the field has responded with Bayesian, conformal, and ensemble approaches that make this feasible. Third, digital patient representations—the “cancer digital twin”—have moved from speculative concept to early clinical implementation, demonstrated by prospective pilot studies at leading cancer centers [9, 85, 93].
12.2 Key Findings of This Review
This review synthesizes over 110 seminal papers to document the following principal findings:
On modeling foundations: Stochastic and spatially resolved models (Eqs. 3–4) outperform classical deterministic formulations in capturing resistance emergence and microenvironmental heterogeneity, but at substantially higher data and computational costs. The Gompertz and logistic formulations remain the appropriate baseline for clinical growth monitoring precisely because their interpretability and low parameter burden are assets in data-sparse settings.
On therapy modeling: PK/PD optimization of chemotherapy (Eqs. 1–2 in Section 3) has demonstrated measurable clinical benefit in reducing adverse events while maintaining efficacy, with multi-objective optimization frameworks now achieving this balance algorithmically rather than empirically. Radiotherapy modeling is undergoing a crisis of confidence in the linear-quadratic model at high doses per fraction, demanding new radiobiological formalisms for SBRT and FLASH modalities.
On immunotherapy and CAR-T: The tumor-immune microenvironment is the most mathematically undercharacterized domain in the field. Published CAR-T models capture expansion-contraction kinetics and antigen-dependent killing, but antigen heterogeneity, spatial barriers, and exhaustion dynamics remain inadequately formalized. Optimal control frameworks (Section 4.1) offer a principled path to dosing optimization but require prospective clinical trial validation.
On AI: Deep learning achieves superior predictive performance when n exceeds approximately 10^3, but the bias-variance decomposition (Eq. 5) establishes that mechanistic models dominate in the data-sparse regimes prevalent in rare cancers and early-phase trials. The identified AI risks—spurious correlation, demographic bias, distribution shift, and automation bias—are not hypothetical: they have been documented empirically and must be systematically addressed before clinical deployment. Conformal prediction (Eq. 22) provides the most robust uncertainty quantification framework currently available.
On clinical translation: The Clinical Readiness Level (CRL) assessment in Table 3 reveals a striking gradient: while PK/PD and radiotherapy models operate at TRL 7–9, the most sophisticated hybrid and digital twin approaches remain at TRL 3–4. This eight-level gap represents not a failure of the science but a reminder that clinical translation requires evidence of prospective validity, regulatory frameworks, and implementation infrastructure that the field is only beginning to build.
12.3 The Central Challenges Remain Structural
The review’s critical analysis (Section 7) reveals that the principal barriers to clinical impact are structural rather than technical. They will not be resolved by incremental model refinement alone. Three challenges are foundational:
The data-model mismatch: Clinical data collection is designed around patient care, not model calibration. The result is chronically sparse, irregularly sampled, and heterogeneously measured data that creates non-identifiability in mechanistic models and underpowered training sets for deep learning. Federated learning (Section 9.1, Eq. 9) and active learning protocols that coordinate model-driven measurement timing with clinical workflows represent the most practical near-term paths forward.
The validation gap: Internal model validation remains the norm; prospective, multi-institutional external validation the exception. The reproducibility crisis documented in Section 7.6 will not self-correct: it requires structural incentives, including pre-registration of modeling studies, mandatory external validation cohorts in funding requirements, and open benchmark datasets with agreed evaluation protocols. The proposed “Model Hub for Oncology” (Section 9.5) addresses this infrastructure deficit directly.
The trust deficit: Clinicians do not yet have the tools, training, or institutional frameworks to evaluate, audit, and appropriately trust or distrust mathematical model outputs. This is not a failure of clinical judgment—it is a failure of the field to produce models whose uncertainty is legible, whose failure modes are documented, and whose outputs are integrated into workflows rather than imposed upon them. The five-point AI safety principle articulated in Section 7.10 represents a minimum standard, not a ceiling.
12.4 The Path Forward: Synthesis of Evidence
The evidence synthesized in this review points toward a convergent architectural vision for the next generation of clinical tumor treatment models. That vision—embodied conceptually in the Causal-Physical Neural Network (CPNN) framework proposed in Section 10—has four defining characteristics:
Causal grounding: Models must encode known biological mechanisms as structural constraints rather than learnable correlations. This means embedding pathway knowledge into model architecture, not merely using it as motivation for feature selection. Structural causal models (Eq. 19) provide the formal language for this commitment.
Physics-constrained learning: Neural components must respect conservation laws and biological positivity constraints by construction, not merely as soft penalties. The Neural ODE formulation (Eq. 20) with architectural positivity guarantees represents the current best practice; future work should extend this to spatiotemporal PDE settings.
Calibrated uncertainty at every output: No clinical prediction without a calibrated interval. Conformal prediction (Eq. 22) achieves this without parametric assumptions; it should be considered a minimum requirement for any model targeting treatment decision support.
Continuous personalization from sparse data: Models must be designed to update from the 2–5 measurements realistic in clinical practice, not the 50+ required by current fine-tuning approaches. The variational encoder in CPNN Module 3 (Eqs. 21a–b) provides a principled architecture for this low-data personalization regime.
These four characteristics are not aspirational. Each has been demonstrated in isolation in published literature [17, 62, 75, 84, 89, 91]. The scientific contribution required is their integration into a validated, clinically deployable system—a challenge that is fundamentally one of execution, collaboration, and institutional will as much as one of technical innovation.
12.5 A Final Word on Identity and Responsibility
Mathematical oncology stands at an inflection point. The field has demonstrated, beyond reasonable doubt, that quantitative models can predict tumor behavior, optimize treatment schedules, and identify resistance mechanisms with accuracy that exceeds unaided clinical intuition in controlled settings. The question is no longer whether mathematical models have a role in oncology—they demonstrably do. The question is whether the field will take seriously the full responsibility that clinical ambition entails.
That responsibility includes honest reporting of model limitations alongside successes; prospective validation before, not after, clinical deployment; equity audits as a standard component of model development; and genuine partnership with clinicians, patients, and regulators in defining what “useful” and “safe” mean in this context. The mathematicians and computer scientists who build these models, and the oncologists who might one day use them, share a common obligation: to ensure that the sophistication of the tools matches the seriousness of the purpose they serve.
The past five years have established the foundation. The next five will determine whether mathematical oncology fulfills its promise as a pillar of personalized cancer care, or remains a scientifically impressive but clinically peripheral enterprise. This review documents the evidence, maps the gaps, and proposes a path. The work of traversing that path remains ahead.
Clearly Auctoresonline and particularly Psychology and Mental Health Care Journal is dedicated to improving health care services for individuals and populations. The editorial boards' ability to efficiently recognize and share the global importance of health literacy with a variety of stakeholders. Auctoresonline publishing platform can be used to facilitate of optimal client-based services and should be added to health care professionals' repertoire of evidence-based health care resources.

Journal of Clinical Cardiology and Cardiovascular Intervention The submission and review process was adequate. However I think that the publication total value should have been enlightened in early fases. Thank you for all.

Journal of Women Health Care and Issues By the present mail, I want to say thank to you and tour colleagues for facilitating my published article. Specially thank you for the peer review process, support from the editorial office. I appreciate positively the quality of your journal.
Journal of Clinical Research and Reports I would be very delighted to submit my testimonial regarding the reviewer board and the editorial office. The reviewer board were accurate and helpful regarding any modifications for my manuscript. And the editorial office were very helpful and supportive in contacting and monitoring with any update and offering help. It was my pleasure to contribute with your promising Journal and I am looking forward for more collaboration.

We would like to thank the Journal of Thoracic Disease and Cardiothoracic Surgery because of the services they provided us for our articles. The peer-review process was done in a very excellent time manner, and the opinions of the reviewers helped us to improve our manuscript further. The editorial office had an outstanding correspondence with us and guided us in many ways. During a hard time of the pandemic that is affecting every one of us tremendously, the editorial office helped us make everything easier for publishing scientific work. Hope for a more scientific relationship with your Journal.

The peer-review process which consisted high quality queries on the paper. I did answer six reviewers’ questions and comments before the paper was accepted. The support from the editorial office is excellent.

Journal of Neuroscience and Neurological Surgery. I had the experience of publishing a research article recently. The whole process was simple from submission to publication. The reviewers made specific and valuable recommendations and corrections that improved the quality of my publication. I strongly recommend this Journal.

Dr. Katarzyna Byczkowska My testimonial covering: "The peer review process is quick and effective. The support from the editorial office is very professional and friendly. Quality of the Clinical Cardiology and Cardiovascular Interventions is scientific and publishes ground-breaking research on cardiology that is useful for other professionals in the field.

Thank you most sincerely, with regard to the support you have given in relation to the reviewing process and the processing of my article entitled "Large Cell Neuroendocrine Carcinoma of The Prostate Gland: A Review and Update" for publication in your esteemed Journal, Journal of Cancer Research and Cellular Therapeutics". The editorial team has been very supportive.

Testimony of Journal of Clinical Otorhinolaryngology: work with your Reviews has been a educational and constructive experience. The editorial office were very helpful and supportive. It was a pleasure to contribute to your Journal.

Dr. Bernard Terkimbi Utoo, I am happy to publish my scientific work in Journal of Women Health Care and Issues (JWHCI). The manuscript submission was seamless and peer review process was top notch. I was amazed that 4 reviewers worked on the manuscript which made it a highly technical, standard and excellent quality paper. I appreciate the format and consideration for the APC as well as the speed of publication. It is my pleasure to continue with this scientific relationship with the esteem JWHCI.

This is an acknowledgment for peer reviewers, editorial board of Journal of Clinical Research and Reports. They show a lot of consideration for us as publishers for our research article “Evaluation of the different factors associated with side effects of COVID-19 vaccination on medical students, Mutah university, Al-Karak, Jordan”, in a very professional and easy way. This journal is one of outstanding medical journal.
Dear Hao Jiang, to Journal of Nutrition and Food Processing We greatly appreciate the efficient, professional and rapid processing of our paper by your team. If there is anything else we should do, please do not hesitate to let us know. On behalf of my co-authors, we would like to express our great appreciation to editor and reviewers.

As an author who has recently published in the journal "Brain and Neurological Disorders". I am delighted to provide a testimonial on the peer review process, editorial office support, and the overall quality of the journal. The peer review process at Brain and Neurological Disorders is rigorous and meticulous, ensuring that only high-quality, evidence-based research is published. The reviewers are experts in their fields, and their comments and suggestions were constructive and helped improve the quality of my manuscript. The review process was timely and efficient, with clear communication from the editorial office at each stage. The support from the editorial office was exceptional throughout the entire process. The editorial staff was responsive, professional, and always willing to help. They provided valuable guidance on formatting, structure, and ethical considerations, making the submission process seamless. Moreover, they kept me informed about the status of my manuscript and provided timely updates, which made the process less stressful. The journal Brain and Neurological Disorders is of the highest quality, with a strong focus on publishing cutting-edge research in the field of neurology. The articles published in this journal are well-researched, rigorously peer-reviewed, and written by experts in the field. The journal maintains high standards, ensuring that readers are provided with the most up-to-date and reliable information on brain and neurological disorders. In conclusion, I had a wonderful experience publishing in Brain and Neurological Disorders. The peer review process was thorough, the editorial office provided exceptional support, and the journal's quality is second to none. I would highly recommend this journal to any researcher working in the field of neurology and brain disorders.

Dear Agrippa Hilda, Journal of Neuroscience and Neurological Surgery, Editorial Coordinator, I trust this message finds you well. I want to extend my appreciation for considering my article for publication in your esteemed journal. I am pleased to provide a testimonial regarding the peer review process and the support received from your editorial office. The peer review process for my paper was carried out in a highly professional and thorough manner. The feedback and comments provided by the authors were constructive and very useful in improving the quality of the manuscript. This rigorous assessment process undoubtedly contributes to the high standards maintained by your journal.

International Journal of Clinical Case Reports and Reviews. I strongly recommend to consider submitting your work to this high-quality journal. The support and availability of the Editorial staff is outstanding and the review process was both efficient and rigorous.

Thank you very much for publishing my Research Article titled “Comparing Treatment Outcome Of Allergic Rhinitis Patients After Using Fluticasone Nasal Spray And Nasal Douching" in the Journal of Clinical Otorhinolaryngology. As Medical Professionals we are immensely benefited from study of various informative Articles and Papers published in this high quality Journal. I look forward to enriching my knowledge by regular study of the Journal and contribute my future work in the field of ENT through the Journal for use by the medical fraternity. The support from the Editorial office was excellent and very prompt. I also welcome the comments received from the readers of my Research Article.

Dear Erica Kelsey, Editorial Coordinator of Cancer Research and Cellular Therapeutics Our team is very satisfied with the processing of our paper by your journal. That was fast, efficient, rigorous, but without unnecessary complications. We appreciated the very short time between the submission of the paper and its publication on line on your site.

I am very glad to say that the peer review process is very successful and fast and support from the Editorial Office. Therefore, I would like to continue our scientific relationship for a long time. And I especially thank you for your kindly attention towards my article. Have a good day!

"We recently published an article entitled “Influence of beta-Cyclodextrins upon the Degradation of Carbofuran Derivatives under Alkaline Conditions" in the Journal of “Pesticides and Biofertilizers” to show that the cyclodextrins protect the carbamates increasing their half-life time in the presence of basic conditions This will be very helpful to understand carbofuran behaviour in the analytical, agro-environmental and food areas. We greatly appreciated the interaction with the editor and the editorial team; we were particularly well accompanied during the course of the revision process, since all various steps towards publication were short and without delay".

I would like to express my gratitude towards you process of article review and submission. I found this to be very fair and expedient. Your follow up has been excellent. I have many publications in national and international journal and your process has been one of the best so far. Keep up the great work.

We are grateful for this opportunity to provide a glowing recommendation to the Journal of Psychiatry and Psychotherapy. We found that the editorial team were very supportive, helpful, kept us abreast of timelines and over all very professional in nature. The peer review process was rigorous, efficient and constructive that really enhanced our article submission. The experience with this journal remains one of our best ever and we look forward to providing future submissions in the near future.

I am very pleased to serve as EBM of the journal, I hope many years of my experience in stem cells can help the journal from one way or another. As we know, stem cells hold great potential for regenerative medicine, which are mostly used to promote the repair response of diseased, dysfunctional or injured tissue using stem cells or their derivatives. I think Stem Cell Research and Therapeutics International is a great platform to publish and share the understanding towards the biology and translational or clinical application of stem cells.

I would like to give my testimony in the support I have got by the peer review process and to support the editorial office where they were of asset to support young author like me to be encouraged to publish their work in your respected journal and globalize and share knowledge across the globe. I really give my great gratitude to your journal and the peer review including the editorial office.

I am delighted to publish our manuscript entitled "A Perspective on Cocaine Induced Stroke - Its Mechanisms and Management" in the Journal of Neuroscience and Neurological Surgery. The peer review process, support from the editorial office, and quality of the journal are excellent. The manuscripts published are of high quality and of excellent scientific value. I recommend this journal very much to colleagues.

Dr.Tania Muñoz, My experience as researcher and author of a review article in The Journal Clinical Cardiology and Interventions has been very enriching and stimulating. The editorial team is excellent, performs its work with absolute responsibility and delivery. They are proactive, dynamic and receptive to all proposals. Supporting at all times the vast universe of authors who choose them as an option for publication. The team of review specialists, members of the editorial board, are brilliant professionals, with remarkable performance in medical research and scientific methodology. Together they form a frontline team that consolidates the JCCI as a magnificent option for the publication and review of high-level medical articles and broad collective interest. I am honored to be able to share my review article and open to receive all your comments.

“The peer review process of JPMHC is quick and effective. Authors are benefited by good and professional reviewers with huge experience in the field of psychology and mental health. The support from the editorial office is very professional. People to contact to are friendly and happy to help and assist any query authors might have. Quality of the Journal is scientific and publishes ground-breaking research on mental health that is useful for other professionals in the field”.

Dear editorial department: On behalf of our team, I hereby certify the reliability and superiority of the International Journal of Clinical Case Reports and Reviews in the peer review process, editorial support, and journal quality. Firstly, the peer review process of the International Journal of Clinical Case Reports and Reviews is rigorous, fair, transparent, fast, and of high quality. The editorial department invites experts from relevant fields as anonymous reviewers to review all submitted manuscripts. These experts have rich academic backgrounds and experience, and can accurately evaluate the academic quality, originality, and suitability of manuscripts. The editorial department is committed to ensuring the rigor of the peer review process, while also making every effort to ensure a fast review cycle to meet the needs of authors and the academic community. Secondly, the editorial team of the International Journal of Clinical Case Reports and Reviews is composed of a group of senior scholars and professionals with rich experience and professional knowledge in related fields. The editorial department is committed to assisting authors in improving their manuscripts, ensuring their academic accuracy, clarity, and completeness. Editors actively collaborate with authors, providing useful suggestions and feedback to promote the improvement and development of the manuscript. We believe that the support of the editorial department is one of the key factors in ensuring the quality of the journal. Finally, the International Journal of Clinical Case Reports and Reviews is renowned for its high- quality articles and strict academic standards. The editorial department is committed to publishing innovative and academically valuable research results to promote the development and progress of related fields. The International Journal of Clinical Case Reports and Reviews is reasonably priced and ensures excellent service and quality ratio, allowing authors to obtain high-level academic publishing opportunities in an affordable manner. I hereby solemnly declare that the International Journal of Clinical Case Reports and Reviews has a high level of credibility and superiority in terms of peer review process, editorial support, reasonable fees, and journal quality. Sincerely, Rui Tao.

Clinical Cardiology and Cardiovascular Interventions I testity the covering of the peer review process, support from the editorial office, and quality of the journal.

Clinical Cardiology and Cardiovascular Interventions, we deeply appreciate the interest shown in our work and its publication. It has been a true pleasure to collaborate with you. The peer review process, as well as the support provided by the editorial office, have been exceptional, and the quality of the journal is very high, which was a determining factor in our decision to publish with you.
The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews journal clinically in the future time.

Clinical Cardiology and Cardiovascular Interventions, I would like to express my sincerest gratitude for the trust placed in our team for the publication in your journal. It has been a true pleasure to collaborate with you on this project. I am pleased to inform you that both the peer review process and the attention from the editorial coordination have been excellent. Your team has worked with dedication and professionalism to ensure that your publication meets the highest standards of quality. We are confident that this collaboration will result in mutual success, and we are eager to see the fruits of this shared effort.

Dear Dr. Jessica Magne, Editorial Coordinator 0f Clinical Cardiology and Cardiovascular Interventions, I hope this message finds you well. I want to express my utmost gratitude for your excellent work and for the dedication and speed in the publication process of my article titled "Navigating Innovation: Qualitative Insights on Using Technology for Health Education in Acute Coronary Syndrome Patients." I am very satisfied with the peer review process, the support from the editorial office, and the quality of the journal. I hope we can maintain our scientific relationship in the long term.
Dear Monica Gissare, - Editorial Coordinator of Nutrition and Food Processing. ¨My testimony with you is truly professional, with a positive response regarding the follow-up of the article and its review, you took into account my qualities and the importance of the topic¨.

Dear Editorial Coordinator of the Journal of Nutrition and Food Processing! "I would like to thank the Journal of Nutrition and Food Processing for including and publishing my article. The peer review process was very quick, movement and precise. The Editorial Board has done an extremely conscientious job with much help, valuable comments and advices. I find the journal very valuable from a professional point of view, thank you very much for allowing me to be part of it and I would like to participate in the future!”

Dealing with The Journal of Neurology and Neurological Surgery was very smooth and comprehensive. The office staff took time to address my needs and the response from editors and the office was prompt and fair. I certainly hope to publish with this journal again.Their professionalism is apparent and more than satisfactory. Susan Weiner

My Testimonial Covering as fellowing: Lin-Show Chin. The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews.

My experience publishing in Psychology and Mental Health Care was exceptional. The peer review process was rigorous and constructive, with reviewers providing valuable insights that helped enhance the quality of our work. The editorial team was highly supportive and responsive, making the submission process smooth and efficient. The journal's commitment to high standards and academic rigor makes it a respected platform for quality research. I am grateful for the opportunity to publish in such a reputable journal.
My experience publishing in International Journal of Clinical Case Reports and Reviews was exceptional. I Come forth to Provide a Testimonial Covering the Peer Review Process and the editorial office for the Professional and Impartial Evaluation of the Manuscript.

I would like to offer my testimony in the support. I have received through the peer review process and support the editorial office where they are to support young authors like me, encourage them to publish their work in your esteemed journals, and globalize and share knowledge globally. I really appreciate your journal, peer review, and editorial office.
Dear Agrippa Hilda- Editorial Coordinator of Journal of Neuroscience and Neurological Surgery, "The peer review process was very quick and of high quality, which can also be seen in the articles in the journal. The collaboration with the editorial office was very good."

I would like to express my sincere gratitude for the support and efficiency provided by the editorial office throughout the publication process of my article, “Delayed Vulvar Metastases from Rectal Carcinoma: A Case Report.” I greatly appreciate the assistance and guidance I received from your team, which made the entire process smooth and efficient. The peer review process was thorough and constructive, contributing to the overall quality of the final article. I am very grateful for the high level of professionalism and commitment shown by the editorial staff, and I look forward to maintaining a long-term collaboration with the International Journal of Clinical Case Reports and Reviews.
To Dear Erin Aust, I would like to express my heartfelt appreciation for the opportunity to have my work published in this esteemed journal. The entire publication process was smooth and well-organized, and I am extremely satisfied with the final result. The Editorial Team demonstrated the utmost professionalism, providing prompt and insightful feedback throughout the review process. Their clear communication and constructive suggestions were invaluable in enhancing my manuscript, and their meticulous attention to detail and dedication to quality are truly commendable. Additionally, the support from the Editorial Office was exceptional. From the initial submission to the final publication, I was guided through every step of the process with great care and professionalism. The team's responsiveness and assistance made the entire experience both easy and stress-free. I am also deeply impressed by the quality and reputation of the journal. It is an honor to have my research featured in such a respected publication, and I am confident that it will make a meaningful contribution to the field.

"I am grateful for the opportunity of contributing to [International Journal of Clinical Case Reports and Reviews] and for the rigorous review process that enhances the quality of research published in your esteemed journal. I sincerely appreciate the time and effort of your team who have dedicatedly helped me in improvising changes and modifying my manuscript. The insightful comments and constructive feedback provided have been invaluable in refining and strengthening my work".

I thank the ‘Journal of Clinical Research and Reports’ for accepting this article for publication. This is a rigorously peer reviewed journal which is on all major global scientific data bases. I note the review process was prompt, thorough and professionally critical. It gave us an insight into a number of important scientific/statistical issues. The review prompted us to review the relevant literature again and look at the limitations of the study. The peer reviewers were open, clear in the instructions and the editorial team was very prompt in their communication. This journal certainly publishes quality research articles. I would recommend the journal for any future publications.

Dear Jessica Magne, with gratitude for the joint work. Fast process of receiving and processing the submitted scientific materials in “Clinical Cardiology and Cardiovascular Interventions”. High level of competence of the editors with clear and correct recommendations and ideas for enriching the article.

We found the peer review process quick and positive in its input. The support from the editorial officer has been very agile, always with the intention of improving the article and taking into account our subsequent corrections.

My article, titled 'No Way Out of the Smartphone Epidemic Without Considering the Insights of Brain Research,' has been republished in the International Journal of Clinical Case Reports and Reviews. The review process was seamless and professional, with the editors being both friendly and supportive. I am deeply grateful for their efforts.
To Dear Erin Aust – Editorial Coordinator of Journal of General Medicine and Clinical Practice! I declare that I am absolutely satisfied with your work carried out with great competence in following the manuscript during the various stages from its receipt, during the revision process to the final acceptance for publication. Thank Prof. Elvira Farina

Dear Jessica, and the super professional team of the ‘Clinical Cardiology and Cardiovascular Interventions’ I am sincerely grateful to the coordinated work of the journal team for the no problem with the submission of my manuscript: “Cardiometabolic Disorders in A Pregnant Woman with Severe Preeclampsia on the Background of Morbid Obesity (Case Report).” The review process by 5 experts was fast, and the comments were professional, which made it more specific and academic, and the process of publication and presentation of the article was excellent. I recommend that my colleagues publish articles in this journal, and I am interested in further scientific cooperation. Sincerely and best wishes, Dr. Oleg Golyanovskiy.

Dear Ashley Rosa, Editorial Coordinator of the journal - Psychology and Mental Health Care. " The process of obtaining publication of my article in the Psychology and Mental Health Journal was positive in all areas. The peer review process resulted in a number of valuable comments, the editorial process was collaborative and timely, and the quality of this journal has been quickly noticed, resulting in alternative journals contacting me to publish with them." Warm regards, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. I appreciate the journal (JCCI) editorial office support, the entire team leads were always ready to help, not only on technical front but also on thorough process. Also, I should thank dear reviewers’ attention to detail and creative approach to teach me and bring new insights by their comments. Surely, more discussions and introduction of other hemodynamic devices would provide better prevention and management of shock states. Your efforts and dedication in presenting educational materials in this journal are commendable. Best wishes from, Farahnaz Fallahian.
Dear Maria Emerson, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. I am delighted to have published our manuscript, "Acute Colonic Pseudo-Obstruction (ACPO): A rare but serious complication following caesarean section." I want to thank the editorial team, especially Maria Emerson, for their prompt review of the manuscript, quick responses to queries, and overall support. Yours sincerely Dr. Victor Olagundoye.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. Many thanks for publishing this manuscript after I lost confidence the editors were most helpful, more than other journals Best wishes from, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Agrippa Hilda, Editorial Coordinator, Journal of Neuroscience and Neurological Surgery. The entire process including article submission, review, revision, and publication was extremely easy. The journal editor was prompt and helpful, and the reviewers contributed to the quality of the paper. Thank you so much! Eric Nussbaum, MD
Dr Hala Al Shaikh This is to acknowledge that the peer review process for the article ’ A Novel Gnrh1 Gene Mutation in Four Omani Male Siblings, Presentation and Management ’ sent to the International Journal of Clinical Case Reports and Reviews was quick and smooth. The editorial office was prompt with easy communication.

Dear Erin Aust, Editorial Coordinator, Journal of General Medicine and Clinical Practice. We are pleased to share our experience with the “Journal of General Medicine and Clinical Practice”, following the successful publication of our article. The peer review process was thorough and constructive, helping to improve the clarity and quality of the manuscript. We are especially thankful to Ms. Erin Aust, the Editorial Coordinator, for her prompt communication and continuous support throughout the process. Her professionalism ensured a smooth and efficient publication experience. The journal upholds high editorial standards, and we highly recommend it to fellow researchers seeking a credible platform for their work. Best wishes By, Dr. Rakhi Mishra.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. The peer review process of the journal of Clinical Cardiology and Cardiovascular Interventions was excellent and fast, as was the support of the editorial office and the quality of the journal. Kind regards Walter F. Riesen Prof. Dr. Dr. h.c. Walter F. Riesen.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. Thank you for publishing our article, Exploring Clozapine's Efficacy in Managing Aggression: A Multiple Single-Case Study in Forensic Psychiatry in the international journal of clinical case reports and reviews. We found the peer review process very professional and efficient. The comments were constructive, and the whole process was efficient. On behalf of the co-authors, I would like to thank you for publishing this article. With regards, Dr. Jelle R. Lettinga.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, I would like to express my deep admiration for the exceptional professionalism demonstrated by your journal. I am thoroughly impressed by the speed of the editorial process, the substantive and insightful reviews, and the meticulous preparation of the manuscript for publication. Additionally, I greatly appreciate the courteous and immediate responses from your editorial office to all my inquiries. Best Regards, Dariusz Ziora

Dear Chrystine Mejia, Editorial Coordinator, Journal of Neurodegeneration and Neurorehabilitation, Auctores Publishing LLC, We would like to thank the editorial team for the smooth and high-quality communication leading up to the publication of our article in the Journal of Neurodegeneration and Neurorehabilitation. The reviewers have extensive knowledge in the field, and their relevant questions helped to add value to our publication. Kind regards, Dr. Ravi Shrivastava.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, Auctores Publishing LLC, USA Office: +1-(302)-520-2644. I would like to express my sincere appreciation for the efficient and professional handling of my case report by the ‘Journal of Clinical Case Reports and Studies’. The peer review process was not only fast but also highly constructive—the reviewers’ comments were clear, relevant, and greatly helped me improve the quality and clarity of my manuscript. I also received excellent support from the editorial office throughout the process. Communication was smooth and timely, and I felt well guided at every stage, from submission to publication. The overall quality and rigor of the journal are truly commendable. I am pleased to have published my work with Journal of Clinical Case Reports and Studies, and I look forward to future opportunities for collaboration. Sincerely, Aline Tollet, UCLouvain.

Dear Ms. Mayra Duenas, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. “The International Journal of Clinical Case Reports and Reviews represented the “ideal house” to share with the research community a first experience with the use of the Simeox device for speech rehabilitation. High scientific reputation and attractive website communication were first determinants for the selection of this Journal, and the following submission process exceeded expectations: fast but highly professional peer review, great support by the editorial office, elegant graphic layout. Exactly what a dynamic research team - also composed by allied professionals - needs!" From, Chiara Beccaluva, PT - Italy.

Dear Maria Emerson, Editorial Coordinator, we have deeply appreciated the professionalism demonstrated by the International Journal of Clinical Case Reports and Reviews. The reviewers have extensive knowledge of our field and have been very efficient and fast in supporting the process. I am really looking forward to further collaboration. Thanks. Best regards, Dr. Claudio Ligresti
Dear Chrystine Mejia, Editorial Coordinator, Journal of Neurodegeneration and Neurorehabilitation. “The peer review process was efficient and constructive, and the editorial office provided excellent communication and support throughout. The journal ensures scientific rigor and high editorial standards, while also offering a smooth and timely publication process. We sincerely appreciate the work of the editorial team in facilitating the dissemination of innovative approaches such as the Bonori Method.” Best regards, Dr. Matteo Bonori.

I recommend without hesitation submitting relevant papers on medical decision making to the International Journal of Clinical Case Reports and Reviews. I am very grateful to the editorial staff. Maria Emerson was a pleasure to communicate with. The time from submission to publication was an extremely short 3 weeks. The editorial staff submitted the paper to three reviewers. Two of the reviewers commented positively on the value of publishing the paper. The editorial staff quickly recognized the third reviewer’s comments as an unjust attempt to reject the paper. I revised the paper as recommended by the first two reviewers.

Dear Maria Emerson, Editorial Coordinator, Journal of Clinical Research and Reports. Thank you for publishing our case report: "Clinical Case of Effective Fetal Stem Cells Treatment in a Patient with Autism Spectrum Disorder" within the "Journal of Clinical Research and Reports" being submitted by the team of EmCell doctors from Kyiv, Ukraine. We much appreciate a professional and transparent peer-review process from Auctores. All research Doctors are so grateful to your Editorial Office and Auctores Publishing support! I amiably wish our article publication maintained a top quality of your International Scientific Journal. My best wishes for a prosperity of the Journal of Clinical Research and Reports. Hope our scientific relationship and cooperation will remain long lasting. Thank you very much indeed. Kind regards, Dr. Andriy Sinelnyk Cell Therapy Center EmCell

Dear Editorial Team, Clinical Cardiology and Cardiovascular Interventions. It was truly a rewarding experience to work with the journal “Clinical Cardiology and Cardiovascular Interventions”. The peer review process was insightful and encouraging, helping us refine our work to a higher standard. The editorial office offered exceptional support with prompt and thoughtful communication. I highly value the journal’s role in promoting scientific advancement and am honored to be part of it. Best regards, Meng-Jou Lee, MD, Department of Anesthesiology, National Taiwan University Hospital.

Dear Editorial Team, Journal-Clinical Cardiology and Cardiovascular Interventions, “Publishing my article with Clinical Cardiology and Cardiovascular Interventions has been a highly positive experience. The peer-review process was rigorous yet supportive, offering valuable feedback that strengthened my work. The editorial team demonstrated exceptional professionalism, prompt communication, and a genuine commitment to maintaining the highest scientific standards. I am very pleased with the publication quality and proud to be associated with such a reputable journal.” Warm regards, Dr. Mahmoud Kamal Moustafa Ahmed

Dear Maria Emerson, Editorial Coordinator of ‘International Journal of Clinical Case Reports and Reviews’, I appreciate the opportunity to publish my article with your journal. The editorial office provided clear communication during the submission and review process, and I found the overall experience professional and constructive. Best regards, Elena Salvatore.

Dear Mayra Duenas, Editorial Coordinator of ‘International Journal of Clinical Case Reports and Reviews Herewith I confirm an optimal peer review process and a great support of the editorial office of the present journal
Dear Editorial Team, Clinical Cardiology and Cardiovascular Interventions. I am really grateful for the peers review; their feedback gave me the opportunity to reflect on the message and impact of my work and to ameliorate the article. The editors did a great job in addition by encouraging me to continue with the process of publishing.

Dear Cecilia Lilly, Editorial Coordinator, Endocrinology and Disorders, Thank you so much for your quick response regarding reviewing and all process till publishing our manuscript entitled: Prevalence of Pre-Diabetes and its Associated Risk Factors Among Nile College Students, Sudan. Best regards, Dr Mamoun Magzoub.

International Journal of Clinical Case Reports and Reviews is a high quality journal that has a clear and concise submission process. The peer review process was comprehensive and constructive. Support from the editorial office was excellent, since the administrative staff were responsive. The journal provides a fast and timely publication timeline.

Dear Mayra Duenas, Editorial Coordinator of the journal IJCCR, I write here a little on my experience as an author submitting to the International Journal of Clinical Case Reports and Reviews (IJCCR). This was my first submission to IJCCR and my manuscript was inherently an outsider’s effort. It attempted to broadly identify and then make some sense of life’s under-appreciated mysteries. I initially had responded to a request for possible submissions. I then contacted IJCCR with a tentative topic for a manuscript. They quickly got back with an approval for the submission, but with a particular requirement that it be medically relevant. I then put together a manuscript and submitted it. After the usual back-and-forth over forms and formality, the manuscript was sent off for reviews. Within 2 weeks I got back 4 reviews which were both helpful and also surprising. Surprising in that the topic was somewhat foreign to medical literature. My subsequent updates in response to the reviewer comments went smoothly and in short order I had a series of proofs to evaluate. All in all, the whole publication process seemed outstanding. It was both helpful in terms of the paper’s content and also in terms of its efficient and friendly communications. Thank you all very much. Sincerely, Ted Christopher, Rochester, NY.

Dear Grace Pierce, Editorial Coordinator of the journal IJCCR, I had a very positive experience with Auctores - Journal throughout the publication process. The Editorial Team was highly responsive, professional, and supportive at every stage. I would like to extend my sincere thanks to the Editor: Grace Pierce, for her guidance and assistance. The peer-review process was smooth and constructive, helping improve the quality of my work. I would gladly recommend Auctores Journal to fellow researchers and authors. Dr. SABITA SINHA, Medical Oncologist, MD (Electro Homeopathy).

Dear Maria Emerson, Editorial Coordinator of - Journal of Clinical Research and Reports. ''I am pleased to provide this testimonial following the publication of our recent case report in this journal. The peer review process was rigorous, constructive, thorough, and conducted in a timely manner. The reviewers’ comments were thoughtful, detailed, and highly constructive, contributing substantially to the refinement, clarity, and scientific robustness of our manuscript. The process was conducted with professionalism and academic integrity throughout. The support provided by the editorial office was exemplary. Communication was consistently prompt, clear, and courteous at all stages of the submission and publication process. The editorial team demonstrated a high level of organization and responsiveness, ensuring that all queries were addressed efficiently and that the process remained transparent and well-coordinated. The overall quality of the journal is reflected in its strong editorial standards, commitment to scientific excellence, and dedication to publishing clinically meaningful research. It has been a privilege to publish our work in this journal, and we would welcome the opportunity to contribute further in the future.'' Best wishes from, Dr. Efstratios Trogkanis, Cardiologist.
Dear Reader: We have published several articles in the Auctores Publishing, LLC, journal, Clinical Medical Reviews and Reports in recent years (CMRR). This is an ‘open access’ journal and the following are our observations. From the initial invitation to submit an article, to the final edits of galley proofs, we have found CMRR personnel to be professional, responsive, rapid and thorough. This entire process begins with Catherine Mitchell, Editorial Coordinator. She is simply outstanding, and, I believe, unparalleled in her capacity. I cannot imagine a more responsive and dedicated Editorial Coordinator. As I read the dates and timing of her correspondence with us, it seems that she never sleeps. I hope Auctores Publishing, LLC, appreciates her efforts as much as these authors do. Thank you to Auctores Publishing, LLC, to the Editorial Staff/Board, and to Catherine Mitchell from a grateful author(s).

Dear Maria Emerson, Editorial Coordinator of International Journal of Clinical Case Reports and Reviews, What distinguishes International Journal of Clinical Case Report and Review is not only the scientific rigor of its publications, but the intellectual climate in which research is evaluated. The submission process is refreshingly free of unnecessary formal barriers and bureaucratic rituals that often complicate academic publishing without adding real value. The peer-review system is demanding yet constructive, guided by genuine scientific dialogue rather than hierarchical or authoritarian attitudes. Reviewers act as collaborators in improving the manuscript, not as gatekeepers imposing arbitrary standards. This journal offers a rare balance: high methodological standards combined with a respectful, transparent, and supportive editorial approach. In an era where publishing can feel more burdensome than research itself, this platform restores the original purpose of peer review — to refine ideas, not to obstruct them Prof. Perlat Kapisyzi, FCCP PULMONOLOGIST AND THORACIC IMAGING.
Dear Mercy Grace, Editorial Coordinator of Obstetrics Gynecology and Reproductive Sciences, We would like to express our gratitude for your help at all stages of publishing and editing the article. The editors of the magazine answer all the necessary questions and help at every stage. We will definitely continue to cooperate and publish other works in the Obstetrics Gynecology and Reproductive Sciences! Best wishes from, Alla Konstantinovna Politova,
