AUCTORES
Globalize your Research
Research Article | DOI: https://doi.org/10.31579/2766-2314/096
1 Gujarat University, Ahmedabad, Gujarat, India.
2 Gyan Manjari University, Bhavnagar, Gujarat, India.
*Corresponding Author: Medha Pandya, Gyan Manjari University, Bhavnagar, Gujarat, India.
Citation: Esha Joshi, Medha Pandya, Shilpa Balar, Hiram Saiyed, Saumya Patel, et al, (2023), Pelargonidin-3,5-Diglucoside found in Pomegranate Aril Acts as a Potent HIF-1α Inhibitor in in-Silico Study, J, Biotechnology and Bioprocessing, 4(2); DOI: 10.31579/2766-2314/096
Copyright: © 2023, Medha Pandya. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received: 27 March 2023 | Accepted: 04 April 2023 | Published: 10 April 2023
Keywords: amino acid; neurons; ischemia; omega-3 polyunsaturated acids
Hypoxia-inducible factor 1 alpha (HIF-1α) is a transcription factor that regulates the transcription of over 100 genes involved in erythropoiesis, angiogenesis, autophagy, cell survival, and energy metabolism signaling; it appears to be a potential target in cancer therapy. The aim of this study was to identify natural compounds from Naturally Occurring Plant-based Anti-cancer Compound-Activity-Target (NPACT) database with anticancer potential. Molecular docking of 1574 phytochemicals was carried out using Autodock Vina 1.1.2 using the site-specific docking method and the amino acid residue site was near Asn803 of HIF-1α. The protein-ligand complexes considered for molecular dynamics simulation were HIF1α- Bortezomib (as control), HIF 1α- Pelargonidin-3 ,5-diglucoside and HIF1α- (25S)-5-beta-spirostan-3-beta-ol-3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-glucopyranoside. Furthermore, ADMET evaluation for the best ligands was carried out using the pkCSM ADMET server.
Hypoxia is a decrease in normal tissue oxygen tension and is a universal characteristic of solid tumors; their ability to adapt to hypoxic conditions is critical to their existence and advancement, and hence could be harnessed in cancer therapy as depicted in Figure 1 and it can arise during the avascular state of tumor, or can emerge in established tumors as a result of inadequate angiogenesis that simply delivers insufficient blood supply [1,2,3]. The rules that govern cellular responses to hypoxia are the hypoxia-inducible factors (HIFs)4. In general, higher HIF expression is linked to tumor growth and therapy resistance, which culminates in disease relapse5. HIFs are categorized into three isoforms: HIF-1, HIF-2, and HIF-36. HIF-1 is regarded to be the principal messenger for hypoxia-induced transcriptional responses and is an indiscriminate heterodimeric transcription factor that consists of two subunits: an α and β. The availability of the HIF-1α subunit, which increases under hypoxic environment, is required for HIF-1 function in tumors [3]. Oxygen-dependent post-translational modifications impact the stability, subcellular localization, and transcriptional strength of the α subunit, and thus oxygen concentration [7]. As HIF-1α regulates the transcription of over 100 genes involved in erythropoiesis, angiogenesis, autophagy, cell survival, and energy metabolism signaling; it appears to be a potential target in cancer therapy [8,9]. The main target genes of HIF-1α are LEP, NO, VEGF, LRP1, ADM, TGF-β for angiogenesis; EPO for erythropoiesis; HK1, HK2, GLUT1, GLUT3, LDHA, PKM for metabolism; IGF2, IGF-BP1, IGF-BP2, IGF-BP3, NOS2, TGF-α for cell-survival and C-MYC, ID2, IGF-2, NOS for cell-proliferation [8,9,10,51]. In normoxic conditions, HIF-1α is degraded by two pathways: (i) pVHL dependent pathway and (ii) pVHL independent pathway as shown in Figure 2. pVHL dependent pathway involves acetylation by ARD1 (Arrest Defective 1) on lysine 532 residue, hydroxylation by PHDs (Prolyl-4-Hydroxylases) on proline 402 and 564 residues, followed by acetylation and ubiquitination on lysine 532 residue; hydroxylation and ubiquitination on proline 402 and proline 564 by pVHL (von Hippel-Lindau protein) and ultimately proteosomal degradation of HIF-1α [10]. While the pVHL independent pathway involves hydroxylation on asparagine 803 by FIH-1 (Factor Inhibiting Hypoxia 1) and thereafter binding of FIH-1 with HIF-1α and hence inhibition of HIF1α and P300/CBP (CREB Binding Protein) dimerization [11].
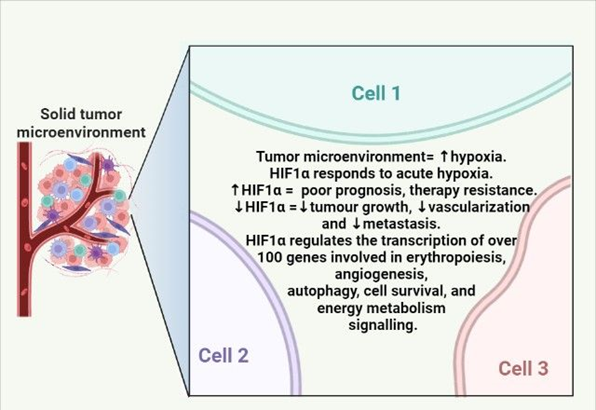
Figure 1: Hypoxia levels are very high in the tumor microenvironment. The above diagram summarizes hypoxia functions in tumor microenvironment (TME).
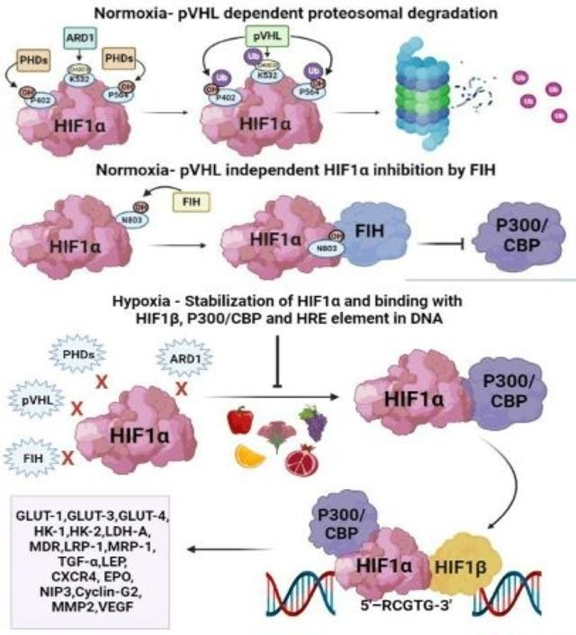
Figure 2: Above figure is modified from Joshi et al. 202351. Degradation and transcription pathway of HIF-1α in normoxia and hypoxia. In normoxic conditions HIF-1α is degraded by several factors including PHDs (PHD1, PHD2 and PHD3), ARD1, pVHL and FIH. However, in hypoxic TME none of these factors are able to degrade HIF-1α and also P300/CBP as well as HIF-1β acts as co-activators for HIF-1α transcription and thus contributes to the upregulation of down-stream genes.
As illustrated in Figure 2, during hypoxic conditions, HIF-1α and P300/CBP dimerization takes place at the asparagine 803 residue which later moves towards the nucleus and binds with HIF-1β and HRE (Hypoxia Response Element) sequence 5’-RCGTG-3’ located on the DNA [10,12].
Despite significant technological breakthroughs in standard therapies, cancer remains the dominant cause of death worldwide. According to the Indian Council of Medical Research, the estimated number of people with cancer is around 2.7 million 2020, each year, new 13.9 lakhs cancer patients are registered and the number of deaths related to Cancer have risen up to 8.5 lakhs in 2020. A fundamental contributing factor to the suboptimal effectiveness of chemotherapy is the unique architectural makeup of tumors. Specifically, in the case of solid tumors, cells situated in hypoxic regions, which are commonly distal from the vasculature, experience constrained access to chemotherapeutic agents due to the atypical vascularization. And also, the MDR1 gene, which is also a target of HIF-1α, represents a significant impediment to effective cancer treatment, as it plays a pivotal role in promoting chemoresistance of cancer cells towards various therapeutic interventions. Chemotherapy is the most efficacious strategy to combat cancer, but it still has detrimental effects. Because of the adverse effects of radiotherapy and chemotherapy, alternative cancer preventive and treatment strategies with no or few side effects are necessary. Natural products can be used as chemotherapeutic and chemo preventive agents as plants and their active constituents have been employed for therapeutic purposes since earlier civilizations. Phytochemicals are currently evolving as gold mines of potent as well as healthier medications against a wide range of life-threatening disorders [13,14,24,26].
Secondary metabolites of plants such as alkaloids possess strong anticancer activity both in-vitro and in-vivo, and can act as CDK or protein kinase inhibitors and can be transformed as novel anticancer agents [16]. A vinca alkaloid, vincristine is a mitotic inhibitor, useful in cancer chemotherapy. Liposomal vincristine has been given FDA approval for treating acute leukemias [17,18]. Polyphenol-derived natural products have gained scientific interest due to their ability to modulate various cancer hallmarks. These hallmarks are a set of distinct biological capabilities acquired by neoplastic cells that enable them to evade host defense mechanisms. Quercetin, a flavonoid, suppresses expression of anti-apoptosis protein Bcl-2 and manages the protein expression of pro-apoptosis Bax and also activates caspase-3, that initiates caspase-3 dependent mitochondrial pathway to induce apoptosis, it also inhibits mTOR phosphorylation and controls AMPK expression in leukemic cells to restrict cell development and trigger cell death [15,24].
Several phytochemicals have exhibited synergistic effects in combination with conventional chemotherapy drugs [19]. For instance, the polyphenol resveratrol, when administered in conjunction with prednisolone, has been observed to decrease the expression of MDR1 protein in cancer cells. Similarly, the flavonoids hesperidin and silibinin, when co-administered with cytarabine at a fixed dose and varied concentration, have been shown to reduce the IC50 value of cytarabine by approximately ~ 5.9 and ~ 4.5 times, respectively. This presents a promising approach for combination therapy with lower cytotoxicity [20]. Furthermore, the phytoestrogen genistein, found in soybeans, has been found to sensitize ovarian cancer cells to the chemotherapy drug cisplatin by inhibiting the NF-κB signaling pathway [21].
Phytochemicals have demonstrated the ability to enhance the bioavailability and uptake of one another. For example, (−)-epicatechin has been observed to increase the incorporation of EGCG in human lung cancer cell line PC-9. Additionally, co-administration of DHA and curcumin at the same concentrations has been shown to significantly augment the uptake of curcumin in human breast cancer SK-BR-3 cells, possibly through alterations in membrane lipid composition [22]. Secondly, in the latest trends of nanotechnology, nanotechnology-mediated delivery of small nucleotides and nucleic acids offers an effective therapeutic strategy for various cancer types [50].
2.1 Homology modelling and structure validation
The quality of HIF-1ɑ was analysed using the PROCHECK program for further structural validation. The graphical representation of the target protein’s predicted tertiary structure and Ramachandran plot analysis are shown in Figure 3. The Ramachandran plot indicates that 63.1% of the residues of the modelled protein were located within the most favored region, 30.5% in additionally allowed region, and 3.4% of the residues in the disallowed region. Thus, the data suggested that the predicted model is reliable for further analysis.

Figure 3: Homology model and Ramachandran plot of Hypoxia inducible factor-1ɑ.
2.2 Analysis of HIF 1 ɑ and its interaction with bortezomib
Recent researchers reported that Bortezomib (PS-341; Velcade, Cambridge, MA) is one of the functional inhibitors of HIF-1ɑ. Bortezomib causes tumor cell death directly and has also been shown to limit tumor adaptation to hypoxia by functionally blocking HIF-1ɑ [23]. The molecular docking of the most potent hypoxic factor HIF-1ɑ was carried out by targeting Asn803 as a docking site for the bortezomib as control. Asn803 serves as a binding site for co-activators P300/CBP required for the transcriptional activation of HIF-1ɑ during hypoxia. In cancer cells, hypoxic microenvironment is a hallmark thus, HIF-1ɑ is stabilized and transcription takes place after the dimerization of HIF-1ɑ with HIF-1β and binding of co-activators P300/CBP and HRE element on the DNA. In-silico molecular docking of HIF-1ɑ leads to -10.8 kcal/mol binding energy with bortezomib as listed in Table 1. The molecular interaction of HIF-1ɑ and bortezomib depicted in Figure 4 and the interacting residues in the cavity are Glu817 with H-bond interaction and Val802, Asn803, Thr796, Tyr798, Leu812, Leu813, Asn689, Val690, Cys800, Glu801 in hydrophobic interactions. Binding of bortezomib with HIF-1ɑ shows that it binds in the co-activators binding pocket as shown in Figure 4: and can inhibit transcriptional activation of HIF-1ɑ.Figure 4: Interaction of Bortezomib in the binding cleft of HIF-1ɑ shown in 3-D representation and 2-D representation (for better clarity) describing ligands interactions by formation of various H-bonds and hydrophobic interactions with protein at the active site of the protein.

Figure 4: Interaction of Bortezomib in the binding cleft of HIF-1ɑ shown in 3-D representation and 2-D representation (for better clarity) describing ligands interactions by formation of various H-bonds and hydrophobic interactions with protein at the active site of the protein.
2.3 Docking of HIF-1ɑ with phytochemicals from NPACT with anti-cancer potential
The NPACT database contains natural chemicals derived from plants that have anticancer properties. Molecular docking study was carried out to evaluate all anticancer natural compounds from this database against HIF-1ɑ. Table 1 lists the top 5 compounds with the lowest binding energies and interacting amino acid residues. The natural flavonoid pelargonidin-3,5-diglucoside reported best binding energy with HIF-1ɑ. Pelargonidin-3,5-diglucoside forming (Figure 5) H-bonds with Val802 (3.04 Ǻ), Ala779 (2.74 Ǻ, 3.30 Ǻ), Gln785 (3.31 Ǻ), Leu783 (3.07 Ǻ), Arg810 (2.89 Ǻ, 2.99 Ǻ), Ser809 (3.01 Ǻ), Asp788 (2.97 Ǻ), Thr798 (2.41 Ǻ, 3.03 Ǻ).

Figure 5: Interaction of Pelargonidin-3,5-diglucoside in the binding cleft of HIF-1ɑ shown in 3-D representation and 2-D representation (for better clarity) describing ligands interactions by formation of various H-bonds and hydrophobic interactions with protein at the active site of the protein.
It forms hydrophilic interactions with Ile607, Met787, Ser786, Pro805, Leu812, Asn803, Tyr798, Leu813, Pro688. Another natural steroid is found in Asparagus officinalis, (25S)-5-beta-spirostan-3-beta-ol-3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-plucopyranoside yields − 11.1 with only one H-bond at Leu797 with bond length 3.15 Ǻ and nineteen hydrophilic bonds at Tyr798, Ala804, Gln814, Leu812, Pro805, Asn803, Ala779, Asp788, Leu783, Arg810, Gln785, Ser809, Ser786, Ile607, Met787, Thr796, Glu817, Val802, Asp83. The third natural saponin named Hederagenin 3-O-beta-D-glucopyranosyl-(1->3)-alpha- L- rhamnopyranosyl-(1->2)-alpha-L- arabinopyranoside binds to HIF-1ɑ with three H-bonds at Asn803 (2.84 Ǻ), Ser786 (3.19 Ǻ), Asp (3.10 Ǻ) and ten hydrophilic bonds Glu817, Tyr798, Ser797, Leu795, Met787, Ser809, Leu812, Pro805, Thr796, Val 802. The binding energies of flavonoid Calyxins K and 7,7''-dimethyllanaraflavone is -10.6 and − 10.5 kcal/mol subsequently and the interacting residues of HIF-1 ɑ with this two-flavonoid depicted in Table1.
| Compound No. | Molecule Name | Binding Energy (kcal/mol) | Class | H- bond Interactions | Hydrophilic Interactions | ||
|---|---|---|---|---|---|---|---|
| No. of Bonds | Residues in Interaction (with bond length) | No. of Bonds | Residues in Interaction (with bond length) | ||||
| Compound | Bortezomib | -10.8 | Anti-cancer Drug | 01 | Glu817 (2.73 Ǻ) | 10 | Val802, Asn803, Thr796, Tyr798, Leu812, Leu813, Asn689, Val690, Cys800, Glu801 |
| 1 | Pelargonidin-3,5-diglucoside | -12 | Flavonoid | 11 | Val802 (3.04 Ǻ), Ala779 (2.74 Ǻ, 3.30 Ǻ), Gln785 (3.31 Ǻ), Leu783 (3.07 Ǻ), Arg810 (2.89 Ǻ, 2.99 Ǻ), Ser809 (3.01 Ǻ), Asp788 (2.97 Ǻ), Thr798 (2.41 Ǻ, 3.03 Ǻ) | 09 | Ile607, Met787, Ser786, Pro805, Leu812, Asn803, Tyr798, Leu813, Pro688 |
| 2 | (25S)-5-beta-spirostan-3-beta-ol 3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-glucopyranoside | -11.2 | Steroid | 01 | Leu797 (3.15 Ǻ) | 19 | Tyr798, Ala804, Gln814, Leu812, Pro805, Asn803, Ala779, Asp788, Leu783, Arg810, Gln785, Ser809, Ser786, Ile607, Met787, Thr796, Glu817, Val802, Asp83 |
| 3 | Hederagenin 3-O-beta-D-glucopyranosyl-(1->3)-alpha-L-rhamnopyranosyl-(1->2)-alpha-L-arabinopyranoside | -11.2 | Saponin | 03 | Asn803 (2.84 Ǻ), Ser786 (3.19 Ǻ), Asp (3.10 Ǻ) | 10 | Glu817, Tyr798, Ser797, Leu795, Met787, Ser809, Leu812, Pro805, Thr796, Val 802 |
| 4 | Calyxins K | -10.6 | Flavonoid | 07 | Cys800 (3.14 Ǻ), Val802 (3.03 Ǻ), Thr796 (2.80 Ǻ), Ser786 (3.02 Ǻ), Ala779 (2.98 Ǻ), Ser797 (2.98 Ǻ), Tyr798 (3.24 Ǻ) | 09 | Glu801, Asn803, Asp788, Ser809, Val690, Leu812, Met787, Pro688, Glu817 |
| 5 | 7,7''-dimethyllanaraflavone | -10.5 | Flavonoid | 03 | Tyr798 (2.89 Ǻ), Ser786 (2.73 Ǻ), Glu817 (3.02 Ǻ) | 12 | Leu812, Ile607, Ser809, Arg810, Pro688, Leu782, Val693, Ile677, Leu783, Met787, Val690, Leu813 |
Table 1: Results of Molecular docking of top molecules of NPACT in interaction with HIF-1α
2.4 Molecular dynamics simulation:
To verify the molecular docking binding affinity results the molecular dynamics simulation approach has been used systematically to investigate the properties of conformational changes that result in changes in protein-ligand interactions, molecular dynamics, and protein folding. MD simulation of complex HIF-1ɑ with Bortezomib and HIF-1ɑ with Pelargonidin-3,5-diglucoside were performed for 100 ns. Analysis of both the complex in context to root mean square deviation (RMSD), root mean square fluctuation (RMSF) and Radius of gyration (Rg) depicted in Figure 6.
After superimposing the receptor on its reference structure, the RMSD of the ligand's heavy atoms was determined. The resulting plots for all the protein-ligand complexes are shown in Figure 6 (a) for bortezomib and 6 (b) for pelargonidin-3,5-diglucoside. This process provides data on how the ligand moves within its binding pocket. The oscillations of bortezomib starts after 0 Ȧ and before 2 Ȧ and goes around 12 Ȧ at 100 ns while for pelargonidin-3,5-diglucoside the highest peak observed is at 16 Ȧ in the middle of 40 ns to 60 ns and that particular point of time, the alterations are not in increased amounts. The pelargonidin-3,5-diglucoside- HIF-1 ɑ complex scale of fluctuation and the divergence between the average RMSD values acclaimed that simulation produced stable trajectories in contrast to control complex.
The functional analysis and the maximum fluctuation in this site are correlated with the root mean square fluctuation analysis (Figure 6c & d). This observed fall in the scores of specific residues indicates that they have a direct role in improving the overall stability of the bound protein complex. The RMSF value is beneficial for illustrating localized minute changes along protein chains and offers information on the dynamic behaviour of proteins in an aqueous simulated environment. The protein districts that change the most during the simulation are shown by the peaks in the diagram. N- and C-terminal protein tails are more prone to change than other regions of the protein. Alpha helices and beta strands, two protein auxiliary regions, are frequently stiffer and more inflexible than unstructured parts. The amino acid residues 350–450 showed high fluctuations in both the complexes. The radius of gyration is calculated and plotted using data collected outside the simulation cell, where each item has its own local coordinate system and no periodic borders are present. The graph for bortezomib starts at 0 ns, reaches the peak at 40 Ȧ in between 30 ns to 40 ns. Pelargonidin-3,5-diglucoside rises at the highest peak above 33.2 Ȧ at 0 ns and falls the lowest to 31.6 Ȧ at 90 ns.

Figure 6: MD simulation trajectory results for Bortezomib (control) and Pelargonidin-3,5-diglucoside. (a) Ligand movement RMSD of Bortezomib (control) after superposing on receptor (b) Ligand movement RMSD of Pelargonidin-3,5-diglucoside (compound 1) after superposing on receptor (c) Solute Protein Nucleic Acid Residue RMSF of control (d) Solute Protein Nucleic Acid Residue RMSF of compound 1 (e)Radius of gyration of solute for Control (f) Radius of gyration of solute for compound 1.
5 .ADMET analysis
The Table 2 summarizes the absorption, distribution, metabolism, excretion and toxicity properties of the top five docked compounds. The compounds that were studied includes- Pelargonidin-3,5-diglucoside (compound 1), (25S)-5-beta-spirostan-3-beta-ol 3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-glucopyranoside (compound 2), Hederagenin 3-O-beta-D-glucopyranosyl-(1->3)-alpha-L-rhamnopyranosyl-(1->2)-alpha-L-arabinopyranoside (compound 3), Calyxins K (compound 4) and 7,7''-dimethyllanaraflavone (compound 5). The main predictor of the absorption properties in in-silico ADMET analysis is the CaCO2 permeability. If a compound's Papp is greater than 8 X 10− 6 cm/s, it is said to have a high CaCO2 permeability. High CaCO2 permeability would produce projected values > 0.90 for the pkCSM predictive model. All the top five compounds have good CaCO2 permeability values as pkCSM predicted. The steady state volume distribution (VDss), for distribution parameters, is the hypothetical volume over which a drug's whole dose would need to be evenly dispersed to have the same concentration as blood plasma. Low VDss is defined as less than 0.71 L/kg (log VDss <-0.15) and high VDss is defined as more than 2.81 L/kg (log VDss > 0.45). Metabolism parameters that are included comprises of- cytochrome P450 inhibitors and CYP2D6/CYP3A4 substrate. Total clearance and renal OCT2 substrate are excretion parameters. Toxicity levels are predicted using ten parameters.
The absorption, distribution, metabolism, excretion, and toxicity properties of the top five docked compounds were summarized in Table 2. The compounds investigated were Pelargonidin-3,5-diglucoside (compound 1), (25S)-5-beta-spirostan-3-beta-ol 3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-glucopyranoside (compound 2), Hederagenin 3-O-beta-D-glucopyranosyl-(1->3)-alpha-L-rhamnopyranosyl-(1->2)-alpha-L-arabinopyranoside (compound 3), Calyxins K (compound 4), and 7,7''-dimethyllanaraflavone (compound 5). In silico ADMET analysis identified CaCO2 permeability as the main predictor of absorption properties. A compound is considered to have high CaCO2 permeability if its Papp value exceeds 8 x 10− 6 cm/s, resulting in projected values > 0.90 for the pkCSM predictive model. All of the top five compounds had good CaCO2 permeability values as predicted by pkCSM. The steady-state volume of distribution (VDss) is a hypothetical volume over which a drug's entire dose must be uniformly distributed to achieve the same concentration as blood plasma. A low VDss is defined as less than 0.71 L/kg (log VDss <-0.15), while a high VDss is defined as more than 2.81 L/kg (log VDss > 0.45) for distribution parameters. Metabolism parameters include cytochrome P450 inhibitors and CYP2D6/CYP3A4 substrates. Total clearance and renal OCT2 substrate are excretion parameters, while toxicity levels are predicted using ten parameters
| Molecule properties | Descriptor | Compound 1 | Compound 2 | Compound 3 | Compound 4 | Compound 5 |
|---|---|---|---|---|---|---|
| Molecular Weight | 595.53 | 913.152 | 913.108 | 582.649 | 566.518 | |
| LogP | -1.8505 | 1.7967 | 1.3453 | 6.8608 | 6.1597 | |
| Rotatable Bonds | 7 | 9 | 9 | 7 | 6 | |
| Acceptors | 14 | 16 | 16 | 8 | 10 | |
| Donors | 10 | 8 | 10 | 4 | 3 | |
| Surface Area | 236.739 | 378.101 | 375.575 | 249.284 | 236.352 | |
| Absorption | Water solubility (log mol/L) | -2.922 | -3.033 | -2.868 | -3.575 | -2.981 |
| CaCO2 permeability (log Papp in 10− 6 cm/s) | 0.827 | 0.882 | 0.543 | 0.824 | 0.207 | |
| Human Intestinal absorption ( Absorbed) | 65.309 | 51.199 | 0 | 94.115 | 87.95 | |
| Skin Permeability (log Kp) | -1.735 | -1.576 | -2.735 | -2.389 | -2.735 | |
| P-glycoprotein substrate (Yes/No) | Yes | Yes | Yes | No | No | |
| P-glycoprotein I inhibitor (Yes/No) | No | Yes | No | Yes | Yes | |
| P-glycoprotein II inhibitor (Yes/No) | No | No | No | Yes | Yes | |
| Distribution | VDss (human) (log L/kg) | -0.462 | -0.4 | -0.433 | -1.244 | -1.128 |
| Fraction unbound (human) (Fu) | 0.207 | 0.431 | 0.394 | 0.334 | 0.423 | |
| BBB permeability (log BB) | -2.503 | -1.826 | -1.671 | -1.313 | -1.824 | |
| CNS permeability (log PS) | -6.134 | -4.689 | -4.89 | -2.711 | -3.475 | |
| Metabolism | CYP2D6 substrate (Yes/No) | No | No | No | No | No |
| CYP3A4 substrate (Yes/No) | No | Yes | No | No | Yes | |
| CYP1A2 inhibitor (Yes/No) | No | No | No | No | No | |
| CYP2C19 inhibitor (Yes/No) | No | No | No | Yes | No | |
| CYP2C9 inhibitor (Yes/No) | No | No | No | Yes | Yes | |
| CYP2D6 inhibitor (Yes/No) | No | No | No | No | No | |
| CYP3A4 inhibitor (Yes/No) | No | No | No | Yes | No | |
| Excretion | Total Clearance (log ml/min/kg) | 0.426 | 0.352 | 0.043 | -0.214 | 0.887 |
| Renal OCT2 substrate (Yes/No) | No | No | No | No | No | |
| Toxicity | AMES toxicity (Yes/No) | No | No | No | No | No |
| Max. tolerated dose (human) (log mg/kg/day) | 0.688 | -2.51 | 0.079 | 0.32 | 0.375 | |
| hERG I inhibitor (Yes/No) | No | No | No | No | No | |
| hERG II inhibitor (Yes/No) | Yes | Yes | No | Yes | Yes | |
| Oral Rat Acute Toxicity (LD50) (mol/kg) | 2.526 | 3.085 | 2.567 | 2.423 | 2.684 | |
| Oral Rat Chronic Toxicity (LOAEL) (log mg/kg bw/day) | 4.453 | 2.845 | 4.93 | 2.027 | 1.957 | |
| Hepatotoxicity | No | No | No | No | No | |
| Skin Sensitisation | No | No | No | No | No | |
| T.Pyriformis toxicity | 0.285 | 0.285 | 0.285 | 0.285 | 0.285 | |
| Minnow toxicity | 9.385 | 7.498 | 6.076 | -1.867 | -3.278 |
HIF-1 ɑ, in normal cells, is tightly regulated and maintains a balance between oxygen supply and demand by promoting angiogenesis, glycolysis, and erythropoiesis. However, in cancer cells, HIF-1ɑ is frequently overexpressed and leads to the activation of various genes that promote tumor growth, angiogenesis, and metastasis [25]. Specifically, HIF-1ɑ promotes glycolysis and the Warburg effect, a metabolic shift in cancer cells that favors glucose metabolism even in the presence of oxygen. This effect provides cancer cells with a survival advantage and allows them to proliferate even in hypoxic environments. HIF-1ɑ is also involved in the regulation of genes that promote angiogenesis [26]. Moreover, HIF-1ɑ has been implicated in the epithelial-to-mesenchymal transition, a process that enhances the invasive and metastatic potential of cancer cells. Overall, the overexpression and dysregulation of HIF-1ɑ in cancer cells play a critical role in tumor progression and aggressiveness, making it an important target for cancer therapy [27]. On the other hand, chemotherapy and standard synthetic drugs can induce several side-effects due to their cytotoxic mechanism of action. For instance, hematological side-effects include bone marrow suppression, resulting in decreased red and white blood cell and platelet counts, which can lead to anemia, increased risk of infections, and bleeding disorders while neurological side-effects include peripheral neuropathy, cognitive dysfunction, and mood changes, which can result from the direct effect of chemotherapy on the nervous system or indirect effects, such as stress and anxiety [28]. On the contrary, the discovery of several other resistance mechanisms has not diminished the significance and prevalence of multidrug resistance (MDR) as the main cause of chemotherapy failure [49].
Hence, this gives rise to the need of treatment using a more natural way. Thus, whether the proposed work can modulate or reduce the activity of HIF-1α using phytochemicals might contribute to the forthwith research on HIF-1ɑ. In the work described above, we molecularly docked HIF-1α with a 1574- natural compound library named NPACT and then considered the compounds with the best docking scores for molecular dynamics (MD) simulations and ADMET analysis. The very first compound, pelargonidin-3,5-diglucoside, with a dock score of -12, is abundant in pomegranate seeds and hence will be easily accessible. The second compound (25S)-5-beta-spirostan-3-beta-ol-3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-Beta -D-Glucopyranoside is found in the Indian Ayurvedic plant Asparagus officinalis or Shatavari and will be a great option for natural cancer treatment.
Pelargonidin-3,5-diglucoside is an anthocyanin, which is a phytochemical that imparts the characteristic red, purple or blue tincture to fruits and vegetables. These pigments are known to exert several health-promoting effects, including anti-cancer properties. High concentrations of Pelargonidin-3,5-diglucoside have been identified in the fruits of Berberis vulgaris (barberry), Punica granatum (pomegranate), Fragaria ananassa (strawberry), as well as in the beans of Phaseolus vulgaris (common bean) and kidney beans [29-31]. Pomegranate juice is recognised to be a significant source of the 3-glucosides and 3,5-diglucosides of pelargonidin, delphinidin and cyanidin [32]. There are reported studies to suggest that pelargonidin-3,5-diglucoside may have anti-cancer activity. In vitro studies have shown that it can inhibit the growth and proliferation of various cancer cell lines, including breast, colon, and liver cancer cells and also shows inhibition of HL-60 cells [33]. It has also been shown to induce apoptosis in these cancer cells, which is a crucial mechanism for preventing cancer growth and spread. While the exact mechanism of action of pelargonidin-3,5-diglucoside in cancer prevention and treatment is not fully understood, it is believed to involve the inhibition of various signaling pathways that are involved in cancer cell growth and survival. Overall, while more research is needed to fully understand the potential anti-cancer effects of pelargonidin-3,5-diglucoside, the available evidence suggests that it may be a promising natural compound for the prevention and treatment of certain types of cancer. In addition, animal studies have demonstrated that pelargonidin-3,5-diglucoside can reduce the incidence and size of tumors in mice with chemically-induced colon cancer.
The second docked compound, (25S)-5-beta-spirostan-3-beta-ol-3-O-alpha-L-rhamnopyranosyl-(1->2)-[alpha-L-rhamnopyranosyl-(1->4)]-beta-D-glucopyranoside with binding energy more than the standard inhibibitor bortezomib, is a plant steroid that can be obtained from the roots of Asparagus officinalis. Pharmacological studies on this plant have revealed its potential therapeutic effects such as anti-inflammatory, cytotoxic, antimutagenic, and antifungal properties. The root extract of Asparagus officinalis has demonstrated cytotoxic activity in various human tumor cell lines such as A2780, Eca-109, HO-8910, CNE, MGC-803, KB, and LTEP-a-2, as well as in the mouse L1210 tumor cell line [34].
Similarly, the next documented phytochemical is a saponin named ranks number three and is found in the roots of Pulsatilla koreana, which were tested for their anticancer effects in vivo using BDF1 mice exhibiting Lewis lung carcinoma (LLC) and it’s in vitro cytotoxic activity against the human solid cancer cell lines SK-MEL-2, A-549, HCT15 and SK-OV-3 [35], whereas the fourth flavonoid, Calyxins K is found in the seeds of Alpinia blepharocalyx and is cytotoxic to murine colon 26-L5 carcinoma and human HT-1080 fibrosarcoma cells [36]. The last flavonoid compound is 7,7''-dimethyllanaraflavone, which is found in leaves of Ouratea hexasperma and Luxemburgia octandra enhanced growth inhibitory action in MCF-7, OVCAR-3 and NCI-H460 cell lines at 3–5 mg/ml (25% control growth) [37].
The molecular dynamics (MD) simulation outcomes indicate that the top two tested compounds exhibit comparable behavior to the FDA-approved drug bortezomib under conditions mimicking those found within the human body, including constant pressure, temperature, and pH. Consequently, these compounds should be considered for further in-vitro investigations. Additionally, the top compounds were subjected to in-silico ADMET analysis through the pkCSM server, revealing that the first and second compounds display potential as inhibitors of HIF-1α. Therefore, these compounds may represent a promising candidate for targeted therapy aimed at HIF-1α inhibition.
5.1 Homology modelling of protein and structure validation:
The amino acid sequence of HIF-1α protein was obtained from the UniProt Database (http://www.uniprot.org/) under the name HUMAN Hypoxia-inducible factor 1-alpha with sequence ID Q16665 and a total sequence length of 826 amino acid residues38. This FASTA-formatted sequence was used for additional investigation. The HIF-1α protein sequence was submitted to the I-TASSER server (https://zhanggroup.org/I-TASSER/) [39,40] for three-dimensional structure prediction using the default settings. For stereo-chemical examination of dihedral angles in modelled protein structure, the predicted models were further evaluated for correct validation and verification using UCLA- SAVES (https://saves.mbi.ucla.edu/) PROCHECK server [39]. PROCHECK examines the overall residue by residue/structural geometry as determined by the Ramachandran plot.
5.2 Molecular Docking:
The 1574 ligands were retrieved from Naturally Occurring Plant-based Anti-cancer Compound-Activity-Target (NPACT) database (https://webs.iiitd.edu.in/raghava/npact/) [41,42]. NPACT focuses solely on anti-cancer natural chemicals found primarily in plants. NPACT is unique in that it provides bioactivities of these natural chemicals against various cancer cell lines as well as their molecular targets. All compounds were optimized and processed in openable tool.
5.3 Molecular Docking and Receptor-Ligand Bond Interactions Analysis:
The homology modelled HIF-1α protein was prepared for docking analysis using MGL tools 1.5.6. The number of grid box points (x × y × z dimensions), center grid box (xyz coordinates) and grid point spacing (Å) for HIF-1α is (24 × 16× 24), 6.667, 65.123, 90.441 and 1Å. The molecular docking of 1574 phytochemicals was carried out in Autodock Vina 1.1.243 using the site-specific docking method and the amino acid residue site was near Asn803 of HIF-1α. Asn803 was selected as a target residue because it allows the binding of HIF-1α to its co-activators CBP/P300 in hypoxic conditions and further promotes transcription. Food and Drugs Administration (FDA) approved HIF-1α inhibitor Bortezomib (PubChem id.: 387447) [44] was taken as control. The binding postures were classified and evaluated based on their binding affinities. To perform protein-ligand binding site analysis, the discovery studio tool was utilized. The best 10 compounds with high binding energy obtained after molecular docking were then analysed for receptor-ligand bond interactions in Biovia Discovery Studio Visualiser [41]. The bond interactions studied included- van der Waals, conventional hydrogen bonds, carbon-hydrogen bonds, alkyl bonds, pi- alkyl bonds, unfavorable acceptor-acceptor, unfavorable donor-donor, pi-anion, pi-sigma, pi-sulphur as well as hydrogen donors and acceptors.
5.4 Molecular dynamics (MD) simulation:
YASARA was used to simulate the molecular dynamics of docked protein-ligand complexes. For MD simulations, the bound protein-ligand complexes HIF1α-Bortezomib (as the control) and HIF1α-Pelargonidin-3,5-diglucoside. In the first instance, they were all energy-minimized to prevent structural instability throughout simulations of dynamics lasting 100 nanoseconds. For simulations employing the YASARA Structure, the AMBER14 force field and TIP3P (transferable intermolecular potential with three points) water model was used. The NPT group was chosen and number of atoms, pressure, and temperature were kept constant. The protein-ligand system was neutralized by adding counter-ions (0.9% NaCl), adjusting the solvent density to 0.997g/l, the pH to 7.4, the temperature to 298 K (Berendsen thermostat), and the pressure to 1 bar (Berendsen barostat). The periodic borders of the simulation cell were turned on, and the steepest gradient energy minimization was run 100 times. Harmonic limitations were adjusted during the equilibration stage. Long-range Coulomb electrostatics was invoked using the particle mesh Ewald approach [45,46].
5.5 ADMET analysis:
Compounds for Absorption, Distribution, Metabolism, Excretion and Toxicity (ADMET) evaluation were selected on the basis of their highest binding energy and molecular dynamics simulation results. ADMET analysis was performed using the pkCSM web-server47,48. pkCSM predicts pharmacokinetic properties of compounds, which relies on graph-based signatures. Smiles format of compounds of interest were retrieved from the PubChem database and were processed for the ADMET results. Thirty predictors were used by pkCSM, and were split into five main classes: absorption (seven predictors), distribution (four predictors), metabolism (seven predictors), excretion (two predictors), and toxicity (ten predictors) [48].
Data Availability Statement:
This article has all the data that was created or analysed during this investigation (and its Supplementary Information files).
EJ is thankful to Scheme of Developing High quality research (SHODH), Education department, Government of Gujarat, India for availing the student support fellowship. All authors would like to thank Department of Zoology, Biomedical Technology, Human Genetics & Wildlife Conservation and Department of Botany, Bioinformatics and Climate Change Impacts Management, School of Sciences at Gujarat University for allowing access to the advance instrumentation and the bioinformatics research facilities. We are also thankful to the Department of Life Sciences, Maharaja Krishna Kumar Sinhji Bhavnagar University, Bhavnagar and also to Department of Biochemistry and Forensic Science, Gujarat University, Ahmedabad, Gujarat.
No specified grant for the above-mentioned research was provided by funding organizations in the public, private, or non-profit sectors.
E.J. and M.P. wrote the manuscript, performed experiments, prepared figures and tables. S.B. helped in preparing the manuscript. U.D., M.P. and S.P. conceptualized the idea, R.M.R., S.P. M.P. and U.D. critically proofread the manuscript.
Competing Interests:
All the authors of the provided manuscript declare no conflicts of interest.
Clearly Auctoresonline and particularly Psychology and Mental Health Care Journal is dedicated to improving health care services for individuals and populations. The editorial boards' ability to efficiently recognize and share the global importance of health literacy with a variety of stakeholders. Auctoresonline publishing platform can be used to facilitate of optimal client-based services and should be added to health care professionals' repertoire of evidence-based health care resources.

Journal of Clinical Cardiology and Cardiovascular Intervention The submission and review process was adequate. However I think that the publication total value should have been enlightened in early fases. Thank you for all.

Journal of Women Health Care and Issues By the present mail, I want to say thank to you and tour colleagues for facilitating my published article. Specially thank you for the peer review process, support from the editorial office. I appreciate positively the quality of your journal.
Journal of Clinical Research and Reports I would be very delighted to submit my testimonial regarding the reviewer board and the editorial office. The reviewer board were accurate and helpful regarding any modifications for my manuscript. And the editorial office were very helpful and supportive in contacting and monitoring with any update and offering help. It was my pleasure to contribute with your promising Journal and I am looking forward for more collaboration.

We would like to thank the Journal of Thoracic Disease and Cardiothoracic Surgery because of the services they provided us for our articles. The peer-review process was done in a very excellent time manner, and the opinions of the reviewers helped us to improve our manuscript further. The editorial office had an outstanding correspondence with us and guided us in many ways. During a hard time of the pandemic that is affecting every one of us tremendously, the editorial office helped us make everything easier for publishing scientific work. Hope for a more scientific relationship with your Journal.

The peer-review process which consisted high quality queries on the paper. I did answer six reviewers’ questions and comments before the paper was accepted. The support from the editorial office is excellent.

Journal of Neuroscience and Neurological Surgery. I had the experience of publishing a research article recently. The whole process was simple from submission to publication. The reviewers made specific and valuable recommendations and corrections that improved the quality of my publication. I strongly recommend this Journal.

Dr. Katarzyna Byczkowska My testimonial covering: "The peer review process is quick and effective. The support from the editorial office is very professional and friendly. Quality of the Clinical Cardiology and Cardiovascular Interventions is scientific and publishes ground-breaking research on cardiology that is useful for other professionals in the field.

Thank you most sincerely, with regard to the support you have given in relation to the reviewing process and the processing of my article entitled "Large Cell Neuroendocrine Carcinoma of The Prostate Gland: A Review and Update" for publication in your esteemed Journal, Journal of Cancer Research and Cellular Therapeutics". The editorial team has been very supportive.

Testimony of Journal of Clinical Otorhinolaryngology: work with your Reviews has been a educational and constructive experience. The editorial office were very helpful and supportive. It was a pleasure to contribute to your Journal.

Dr. Bernard Terkimbi Utoo, I am happy to publish my scientific work in Journal of Women Health Care and Issues (JWHCI). The manuscript submission was seamless and peer review process was top notch. I was amazed that 4 reviewers worked on the manuscript which made it a highly technical, standard and excellent quality paper. I appreciate the format and consideration for the APC as well as the speed of publication. It is my pleasure to continue with this scientific relationship with the esteem JWHCI.

This is an acknowledgment for peer reviewers, editorial board of Journal of Clinical Research and Reports. They show a lot of consideration for us as publishers for our research article “Evaluation of the different factors associated with side effects of COVID-19 vaccination on medical students, Mutah university, Al-Karak, Jordan”, in a very professional and easy way. This journal is one of outstanding medical journal.
Dear Hao Jiang, to Journal of Nutrition and Food Processing We greatly appreciate the efficient, professional and rapid processing of our paper by your team. If there is anything else we should do, please do not hesitate to let us know. On behalf of my co-authors, we would like to express our great appreciation to editor and reviewers.

As an author who has recently published in the journal "Brain and Neurological Disorders". I am delighted to provide a testimonial on the peer review process, editorial office support, and the overall quality of the journal. The peer review process at Brain and Neurological Disorders is rigorous and meticulous, ensuring that only high-quality, evidence-based research is published. The reviewers are experts in their fields, and their comments and suggestions were constructive and helped improve the quality of my manuscript. The review process was timely and efficient, with clear communication from the editorial office at each stage. The support from the editorial office was exceptional throughout the entire process. The editorial staff was responsive, professional, and always willing to help. They provided valuable guidance on formatting, structure, and ethical considerations, making the submission process seamless. Moreover, they kept me informed about the status of my manuscript and provided timely updates, which made the process less stressful. The journal Brain and Neurological Disorders is of the highest quality, with a strong focus on publishing cutting-edge research in the field of neurology. The articles published in this journal are well-researched, rigorously peer-reviewed, and written by experts in the field. The journal maintains high standards, ensuring that readers are provided with the most up-to-date and reliable information on brain and neurological disorders. In conclusion, I had a wonderful experience publishing in Brain and Neurological Disorders. The peer review process was thorough, the editorial office provided exceptional support, and the journal's quality is second to none. I would highly recommend this journal to any researcher working in the field of neurology and brain disorders.

Dear Agrippa Hilda, Journal of Neuroscience and Neurological Surgery, Editorial Coordinator, I trust this message finds you well. I want to extend my appreciation for considering my article for publication in your esteemed journal. I am pleased to provide a testimonial regarding the peer review process and the support received from your editorial office. The peer review process for my paper was carried out in a highly professional and thorough manner. The feedback and comments provided by the authors were constructive and very useful in improving the quality of the manuscript. This rigorous assessment process undoubtedly contributes to the high standards maintained by your journal.

International Journal of Clinical Case Reports and Reviews. I strongly recommend to consider submitting your work to this high-quality journal. The support and availability of the Editorial staff is outstanding and the review process was both efficient and rigorous.

Thank you very much for publishing my Research Article titled “Comparing Treatment Outcome Of Allergic Rhinitis Patients After Using Fluticasone Nasal Spray And Nasal Douching" in the Journal of Clinical Otorhinolaryngology. As Medical Professionals we are immensely benefited from study of various informative Articles and Papers published in this high quality Journal. I look forward to enriching my knowledge by regular study of the Journal and contribute my future work in the field of ENT through the Journal for use by the medical fraternity. The support from the Editorial office was excellent and very prompt. I also welcome the comments received from the readers of my Research Article.

Dear Erica Kelsey, Editorial Coordinator of Cancer Research and Cellular Therapeutics Our team is very satisfied with the processing of our paper by your journal. That was fast, efficient, rigorous, but without unnecessary complications. We appreciated the very short time between the submission of the paper and its publication on line on your site.

I am very glad to say that the peer review process is very successful and fast and support from the Editorial Office. Therefore, I would like to continue our scientific relationship for a long time. And I especially thank you for your kindly attention towards my article. Have a good day!

"We recently published an article entitled “Influence of beta-Cyclodextrins upon the Degradation of Carbofuran Derivatives under Alkaline Conditions" in the Journal of “Pesticides and Biofertilizers” to show that the cyclodextrins protect the carbamates increasing their half-life time in the presence of basic conditions This will be very helpful to understand carbofuran behaviour in the analytical, agro-environmental and food areas. We greatly appreciated the interaction with the editor and the editorial team; we were particularly well accompanied during the course of the revision process, since all various steps towards publication were short and without delay".

I would like to express my gratitude towards you process of article review and submission. I found this to be very fair and expedient. Your follow up has been excellent. I have many publications in national and international journal and your process has been one of the best so far. Keep up the great work.

We are grateful for this opportunity to provide a glowing recommendation to the Journal of Psychiatry and Psychotherapy. We found that the editorial team were very supportive, helpful, kept us abreast of timelines and over all very professional in nature. The peer review process was rigorous, efficient and constructive that really enhanced our article submission. The experience with this journal remains one of our best ever and we look forward to providing future submissions in the near future.

I am very pleased to serve as EBM of the journal, I hope many years of my experience in stem cells can help the journal from one way or another. As we know, stem cells hold great potential for regenerative medicine, which are mostly used to promote the repair response of diseased, dysfunctional or injured tissue using stem cells or their derivatives. I think Stem Cell Research and Therapeutics International is a great platform to publish and share the understanding towards the biology and translational or clinical application of stem cells.

I would like to give my testimony in the support I have got by the peer review process and to support the editorial office where they were of asset to support young author like me to be encouraged to publish their work in your respected journal and globalize and share knowledge across the globe. I really give my great gratitude to your journal and the peer review including the editorial office.

I am delighted to publish our manuscript entitled "A Perspective on Cocaine Induced Stroke - Its Mechanisms and Management" in the Journal of Neuroscience and Neurological Surgery. The peer review process, support from the editorial office, and quality of the journal are excellent. The manuscripts published are of high quality and of excellent scientific value. I recommend this journal very much to colleagues.

Dr.Tania Muñoz, My experience as researcher and author of a review article in The Journal Clinical Cardiology and Interventions has been very enriching and stimulating. The editorial team is excellent, performs its work with absolute responsibility and delivery. They are proactive, dynamic and receptive to all proposals. Supporting at all times the vast universe of authors who choose them as an option for publication. The team of review specialists, members of the editorial board, are brilliant professionals, with remarkable performance in medical research and scientific methodology. Together they form a frontline team that consolidates the JCCI as a magnificent option for the publication and review of high-level medical articles and broad collective interest. I am honored to be able to share my review article and open to receive all your comments.

“The peer review process of JPMHC is quick and effective. Authors are benefited by good and professional reviewers with huge experience in the field of psychology and mental health. The support from the editorial office is very professional. People to contact to are friendly and happy to help and assist any query authors might have. Quality of the Journal is scientific and publishes ground-breaking research on mental health that is useful for other professionals in the field”.

Dear editorial department: On behalf of our team, I hereby certify the reliability and superiority of the International Journal of Clinical Case Reports and Reviews in the peer review process, editorial support, and journal quality. Firstly, the peer review process of the International Journal of Clinical Case Reports and Reviews is rigorous, fair, transparent, fast, and of high quality. The editorial department invites experts from relevant fields as anonymous reviewers to review all submitted manuscripts. These experts have rich academic backgrounds and experience, and can accurately evaluate the academic quality, originality, and suitability of manuscripts. The editorial department is committed to ensuring the rigor of the peer review process, while also making every effort to ensure a fast review cycle to meet the needs of authors and the academic community. Secondly, the editorial team of the International Journal of Clinical Case Reports and Reviews is composed of a group of senior scholars and professionals with rich experience and professional knowledge in related fields. The editorial department is committed to assisting authors in improving their manuscripts, ensuring their academic accuracy, clarity, and completeness. Editors actively collaborate with authors, providing useful suggestions and feedback to promote the improvement and development of the manuscript. We believe that the support of the editorial department is one of the key factors in ensuring the quality of the journal. Finally, the International Journal of Clinical Case Reports and Reviews is renowned for its high- quality articles and strict academic standards. The editorial department is committed to publishing innovative and academically valuable research results to promote the development and progress of related fields. The International Journal of Clinical Case Reports and Reviews is reasonably priced and ensures excellent service and quality ratio, allowing authors to obtain high-level academic publishing opportunities in an affordable manner. I hereby solemnly declare that the International Journal of Clinical Case Reports and Reviews has a high level of credibility and superiority in terms of peer review process, editorial support, reasonable fees, and journal quality. Sincerely, Rui Tao.

Clinical Cardiology and Cardiovascular Interventions I testity the covering of the peer review process, support from the editorial office, and quality of the journal.

Clinical Cardiology and Cardiovascular Interventions, we deeply appreciate the interest shown in our work and its publication. It has been a true pleasure to collaborate with you. The peer review process, as well as the support provided by the editorial office, have been exceptional, and the quality of the journal is very high, which was a determining factor in our decision to publish with you.
The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews journal clinically in the future time.

Clinical Cardiology and Cardiovascular Interventions, I would like to express my sincerest gratitude for the trust placed in our team for the publication in your journal. It has been a true pleasure to collaborate with you on this project. I am pleased to inform you that both the peer review process and the attention from the editorial coordination have been excellent. Your team has worked with dedication and professionalism to ensure that your publication meets the highest standards of quality. We are confident that this collaboration will result in mutual success, and we are eager to see the fruits of this shared effort.

Dear Dr. Jessica Magne, Editorial Coordinator 0f Clinical Cardiology and Cardiovascular Interventions, I hope this message finds you well. I want to express my utmost gratitude for your excellent work and for the dedication and speed in the publication process of my article titled "Navigating Innovation: Qualitative Insights on Using Technology for Health Education in Acute Coronary Syndrome Patients." I am very satisfied with the peer review process, the support from the editorial office, and the quality of the journal. I hope we can maintain our scientific relationship in the long term.
Dear Monica Gissare, - Editorial Coordinator of Nutrition and Food Processing. ¨My testimony with you is truly professional, with a positive response regarding the follow-up of the article and its review, you took into account my qualities and the importance of the topic¨.

Dear Dr. Jessica Magne, Editorial Coordinator 0f Clinical Cardiology and Cardiovascular Interventions, The review process for the article “The Handling of Anti-aggregants and Anticoagulants in the Oncologic Heart Patient Submitted to Surgery” was extremely rigorous and detailed. From the initial submission to the final acceptance, the editorial team at the “Journal of Clinical Cardiology and Cardiovascular Interventions” demonstrated a high level of professionalism and dedication. The reviewers provided constructive and detailed feedback, which was essential for improving the quality of our work. Communication was always clear and efficient, ensuring that all our questions were promptly addressed. The quality of the “Journal of Clinical Cardiology and Cardiovascular Interventions” is undeniable. It is a peer-reviewed, open-access publication dedicated exclusively to disseminating high-quality research in the field of clinical cardiology and cardiovascular interventions. The journal's impact factor is currently under evaluation, and it is indexed in reputable databases, which further reinforces its credibility and relevance in the scientific field. I highly recommend this journal to researchers looking for a reputable platform to publish their studies.
Dear Editorial Coordinator of the Journal of Nutrition and Food Processing! "I would like to thank the Journal of Nutrition and Food Processing for including and publishing my article. The peer review process was very quick, movement and precise. The Editorial Board has done an extremely conscientious job with much help, valuable comments and advices. I find the journal very valuable from a professional point of view, thank you very much for allowing me to be part of it and I would like to participate in the future!”

Dealing with The Journal of Neurology and Neurological Surgery was very smooth and comprehensive. The office staff took time to address my needs and the response from editors and the office was prompt and fair. I certainly hope to publish with this journal again.Their professionalism is apparent and more than satisfactory. Susan Weiner

My Testimonial Covering as fellowing: Lin-Show Chin. The peer reviewers process is quick and effective, the supports from editorial office is excellent, the quality of journal is high. I would like to collabroate with Internatioanl journal of Clinical Case Reports and Reviews.

My experience publishing in Psychology and Mental Health Care was exceptional. The peer review process was rigorous and constructive, with reviewers providing valuable insights that helped enhance the quality of our work. The editorial team was highly supportive and responsive, making the submission process smooth and efficient. The journal's commitment to high standards and academic rigor makes it a respected platform for quality research. I am grateful for the opportunity to publish in such a reputable journal.
My experience publishing in International Journal of Clinical Case Reports and Reviews was exceptional. I Come forth to Provide a Testimonial Covering the Peer Review Process and the editorial office for the Professional and Impartial Evaluation of the Manuscript.

I would like to offer my testimony in the support. I have received through the peer review process and support the editorial office where they are to support young authors like me, encourage them to publish their work in your esteemed journals, and globalize and share knowledge globally. I really appreciate your journal, peer review, and editorial office.
Dear Agrippa Hilda- Editorial Coordinator of Journal of Neuroscience and Neurological Surgery, "The peer review process was very quick and of high quality, which can also be seen in the articles in the journal. The collaboration with the editorial office was very good."

I would like to express my sincere gratitude for the support and efficiency provided by the editorial office throughout the publication process of my article, “Delayed Vulvar Metastases from Rectal Carcinoma: A Case Report.” I greatly appreciate the assistance and guidance I received from your team, which made the entire process smooth and efficient. The peer review process was thorough and constructive, contributing to the overall quality of the final article. I am very grateful for the high level of professionalism and commitment shown by the editorial staff, and I look forward to maintaining a long-term collaboration with the International Journal of Clinical Case Reports and Reviews.
To Dear Erin Aust, I would like to express my heartfelt appreciation for the opportunity to have my work published in this esteemed journal. The entire publication process was smooth and well-organized, and I am extremely satisfied with the final result. The Editorial Team demonstrated the utmost professionalism, providing prompt and insightful feedback throughout the review process. Their clear communication and constructive suggestions were invaluable in enhancing my manuscript, and their meticulous attention to detail and dedication to quality are truly commendable. Additionally, the support from the Editorial Office was exceptional. From the initial submission to the final publication, I was guided through every step of the process with great care and professionalism. The team's responsiveness and assistance made the entire experience both easy and stress-free. I am also deeply impressed by the quality and reputation of the journal. It is an honor to have my research featured in such a respected publication, and I am confident that it will make a meaningful contribution to the field.

"I am grateful for the opportunity of contributing to [International Journal of Clinical Case Reports and Reviews] and for the rigorous review process that enhances the quality of research published in your esteemed journal. I sincerely appreciate the time and effort of your team who have dedicatedly helped me in improvising changes and modifying my manuscript. The insightful comments and constructive feedback provided have been invaluable in refining and strengthening my work".

I thank the ‘Journal of Clinical Research and Reports’ for accepting this article for publication. This is a rigorously peer reviewed journal which is on all major global scientific data bases. I note the review process was prompt, thorough and professionally critical. It gave us an insight into a number of important scientific/statistical issues. The review prompted us to review the relevant literature again and look at the limitations of the study. The peer reviewers were open, clear in the instructions and the editorial team was very prompt in their communication. This journal certainly publishes quality research articles. I would recommend the journal for any future publications.

Dear Jessica Magne, with gratitude for the joint work. Fast process of receiving and processing the submitted scientific materials in “Clinical Cardiology and Cardiovascular Interventions”. High level of competence of the editors with clear and correct recommendations and ideas for enriching the article.

We found the peer review process quick and positive in its input. The support from the editorial officer has been very agile, always with the intention of improving the article and taking into account our subsequent corrections.

My article, titled 'No Way Out of the Smartphone Epidemic Without Considering the Insights of Brain Research,' has been republished in the International Journal of Clinical Case Reports and Reviews. The review process was seamless and professional, with the editors being both friendly and supportive. I am deeply grateful for their efforts.
To Dear Erin Aust – Editorial Coordinator of Journal of General Medicine and Clinical Practice! I declare that I am absolutely satisfied with your work carried out with great competence in following the manuscript during the various stages from its receipt, during the revision process to the final acceptance for publication. Thank Prof. Elvira Farina

Dear Jessica, and the super professional team of the ‘Clinical Cardiology and Cardiovascular Interventions’ I am sincerely grateful to the coordinated work of the journal team for the no problem with the submission of my manuscript: “Cardiometabolic Disorders in A Pregnant Woman with Severe Preeclampsia on the Background of Morbid Obesity (Case Report).” The review process by 5 experts was fast, and the comments were professional, which made it more specific and academic, and the process of publication and presentation of the article was excellent. I recommend that my colleagues publish articles in this journal, and I am interested in further scientific cooperation. Sincerely and best wishes, Dr. Oleg Golyanovskiy.

Dear Ashley Rosa, Editorial Coordinator of the journal - Psychology and Mental Health Care. " The process of obtaining publication of my article in the Psychology and Mental Health Journal was positive in all areas. The peer review process resulted in a number of valuable comments, the editorial process was collaborative and timely, and the quality of this journal has been quickly noticed, resulting in alternative journals contacting me to publish with them." Warm regards, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. I appreciate the journal (JCCI) editorial office support, the entire team leads were always ready to help, not only on technical front but also on thorough process. Also, I should thank dear reviewers’ attention to detail and creative approach to teach me and bring new insights by their comments. Surely, more discussions and introduction of other hemodynamic devices would provide better prevention and management of shock states. Your efforts and dedication in presenting educational materials in this journal are commendable. Best wishes from, Farahnaz Fallahian.
Dear Maria Emerson, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. I am delighted to have published our manuscript, "Acute Colonic Pseudo-Obstruction (ACPO): A rare but serious complication following caesarean section." I want to thank the editorial team, especially Maria Emerson, for their prompt review of the manuscript, quick responses to queries, and overall support. Yours sincerely Dr. Victor Olagundoye.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. Many thanks for publishing this manuscript after I lost confidence the editors were most helpful, more than other journals Best wishes from, Susan Anne Smith, PhD. Australian Breastfeeding Association.

Dear Agrippa Hilda, Editorial Coordinator, Journal of Neuroscience and Neurological Surgery. The entire process including article submission, review, revision, and publication was extremely easy. The journal editor was prompt and helpful, and the reviewers contributed to the quality of the paper. Thank you so much! Eric Nussbaum, MD
Dr Hala Al Shaikh This is to acknowledge that the peer review process for the article ’ A Novel Gnrh1 Gene Mutation in Four Omani Male Siblings, Presentation and Management ’ sent to the International Journal of Clinical Case Reports and Reviews was quick and smooth. The editorial office was prompt with easy communication.

Dear Erin Aust, Editorial Coordinator, Journal of General Medicine and Clinical Practice. We are pleased to share our experience with the “Journal of General Medicine and Clinical Practice”, following the successful publication of our article. The peer review process was thorough and constructive, helping to improve the clarity and quality of the manuscript. We are especially thankful to Ms. Erin Aust, the Editorial Coordinator, for her prompt communication and continuous support throughout the process. Her professionalism ensured a smooth and efficient publication experience. The journal upholds high editorial standards, and we highly recommend it to fellow researchers seeking a credible platform for their work. Best wishes By, Dr. Rakhi Mishra.

Dear Jessica Magne, Editorial Coordinator, Clinical Cardiology and Cardiovascular Interventions, Auctores Publishing LLC. The peer review process of the journal of Clinical Cardiology and Cardiovascular Interventions was excellent and fast, as was the support of the editorial office and the quality of the journal. Kind regards Walter F. Riesen Prof. Dr. Dr. h.c. Walter F. Riesen.

Dear Ashley Rosa, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews, Auctores Publishing LLC. Thank you for publishing our article, Exploring Clozapine's Efficacy in Managing Aggression: A Multiple Single-Case Study in Forensic Psychiatry in the international journal of clinical case reports and reviews. We found the peer review process very professional and efficient. The comments were constructive, and the whole process was efficient. On behalf of the co-authors, I would like to thank you for publishing this article. With regards, Dr. Jelle R. Lettinga.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, I would like to express my deep admiration for the exceptional professionalism demonstrated by your journal. I am thoroughly impressed by the speed of the editorial process, the substantive and insightful reviews, and the meticulous preparation of the manuscript for publication. Additionally, I greatly appreciate the courteous and immediate responses from your editorial office to all my inquiries. Best Regards, Dariusz Ziora

Dear Chrystine Mejia, Editorial Coordinator, Journal of Neurodegeneration and Neurorehabilitation, Auctores Publishing LLC, We would like to thank the editorial team for the smooth and high-quality communication leading up to the publication of our article in the Journal of Neurodegeneration and Neurorehabilitation. The reviewers have extensive knowledge in the field, and their relevant questions helped to add value to our publication. Kind regards, Dr. Ravi Shrivastava.

Dear Clarissa Eric, Editorial Coordinator, Journal of Clinical Case Reports and Studies, Auctores Publishing LLC, USA Office: +1-(302)-520-2644. I would like to express my sincere appreciation for the efficient and professional handling of my case report by the ‘Journal of Clinical Case Reports and Studies’. The peer review process was not only fast but also highly constructive—the reviewers’ comments were clear, relevant, and greatly helped me improve the quality and clarity of my manuscript. I also received excellent support from the editorial office throughout the process. Communication was smooth and timely, and I felt well guided at every stage, from submission to publication. The overall quality and rigor of the journal are truly commendable. I am pleased to have published my work with Journal of Clinical Case Reports and Studies, and I look forward to future opportunities for collaboration. Sincerely, Aline Tollet, UCLouvain.

Dear Ms. Mayra Duenas, Editorial Coordinator, International Journal of Clinical Case Reports and Reviews. “The International Journal of Clinical Case Reports and Reviews represented the “ideal house” to share with the research community a first experience with the use of the Simeox device for speech rehabilitation. High scientific reputation and attractive website communication were first determinants for the selection of this Journal, and the following submission process exceeded expectations: fast but highly professional peer review, great support by the editorial office, elegant graphic layout. Exactly what a dynamic research team - also composed by allied professionals - needs!" From, Chiara Beccaluva, PT - Italy.

Dear Maria Emerson, Editorial Coordinator, we have deeply appreciated the professionalism demonstrated by the International Journal of Clinical Case Reports and Reviews. The reviewers have extensive knowledge of our field and have been very efficient and fast in supporting the process. I am really looking forward to further collaboration. Thanks. Best regards, Dr. Claudio Ligresti